🧬A Primer on AI Protein Design
The field went from predicting what proteins look like to designing ones that have never existed. Here's an intro to how.
Welcome back to Kiin Bio Weekly.
For decades, designing a new protein meant years of directed evolution, rational engineering, and a lot of luck. You started from something nature already made and slowly nudged it toward what you wanted. The success rate was low. The timelines were long.
That’s changing fast. A wave of AI models, led by tools like RFDiffusion, ProteinMPNN, and AlphaFold, has opened up a fundamentally different approach: designing proteins from scratch, computationally, and getting functional molecules on the first try. In the last two years, AI-designed proteins have matched or outperformed naturally evolved ones in binding affinity, stability, and specificity, sometimes by significant margins.
This primer covers what’s actually happening, how the key methods work, and where the field is headed.
Keeping up with AI x life science news can get exhausting.
It’s scattered across LinkedIn, X, Substack, arXiv, Slack, newsletters... and you still somehow miss the things that actually matter. Too much noise, not enough signal.
We’re building something to fix that: a smarter, more powerful way to stay on top of what’s actually relevant to you.
But we want to build it with you, not just for you. Take 2 minutes to tell us what’s missing. What you share will directly shape what we build, and you’ll be the first to benefit from it.
🔬 From Prediction to Design
The story starts with structure prediction. AlphaFold, released by DeepMind in 2020, solved a 50-year-old problem: predicting a protein’s 3D structure from its amino acid sequence. That was transformative for understanding biology, but it didn’t directly design new proteins. It told you what a sequence would fold into, not what sequence would give you the fold you wanted.
The leap to design required inverting the problem. Instead of sequence → structure, the question became: what sequence would fold into a structure that does what I need?
That inversion is what the current generation of tools enables.
🧩 The Key Methods
There are three categories of AI tools driving protein design today, and they work best in combination.
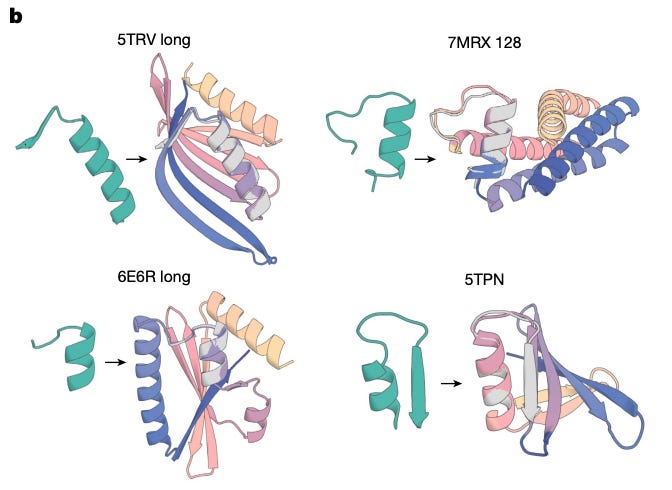
Structure generation creates new protein backbone shapes, the 3D scaffold. The breakthrough here is RFDiffusion, developed by David Baker’s lab at the University of Washington. It uses diffusion models (the same class of generative AI behind image tools like DALL-E) applied to 3D coordinates. You specify what you want, a protein that binds a particular target, wraps around a small molecule, or presents a specific functional site, and the model generates backbone structures that satisfy those constraints. It’s designing architectures that evolution never explored.
Sequence design fills in the amino acid sequence for a given backbone. ProteinMPNN, also from Baker’s lab, takes a 3D structure and predicts which amino acid sequences will fold into it stably. This is the bridge between a computational shape and something you can actually synthesise and test. It recovers native-like sequences with high accuracy and, critically, produces sequences that fold and function when tested experimentally.
Structure prediction closes the loop. AlphaFold (and its open-source successor ESMFold from Meta) validates the designs by predicting whether the designed sequence will actually fold into the intended structure. If the predicted fold matches the designed backbone, confidence is high. If it doesn’t, you iterate.
The typical workflow today: RFDiffusion generates a backbone → ProteinMPNN designs sequences for it → AlphaFold confirms the fold → the best candidates go to the lab.
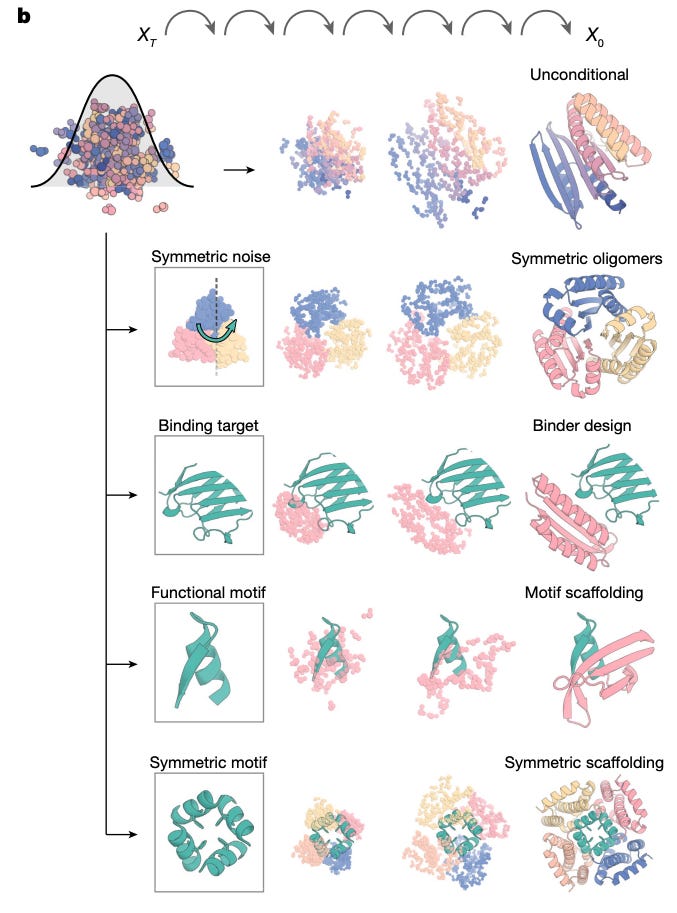
The AI protein design workflow: a diffusion model generates novel backbone structures, a sequence design model fills in amino acids, and a structure prediction model validates the fold before experimental testing.
⚙️ What’s Actually Working
The results from the last 18 months have been striking.
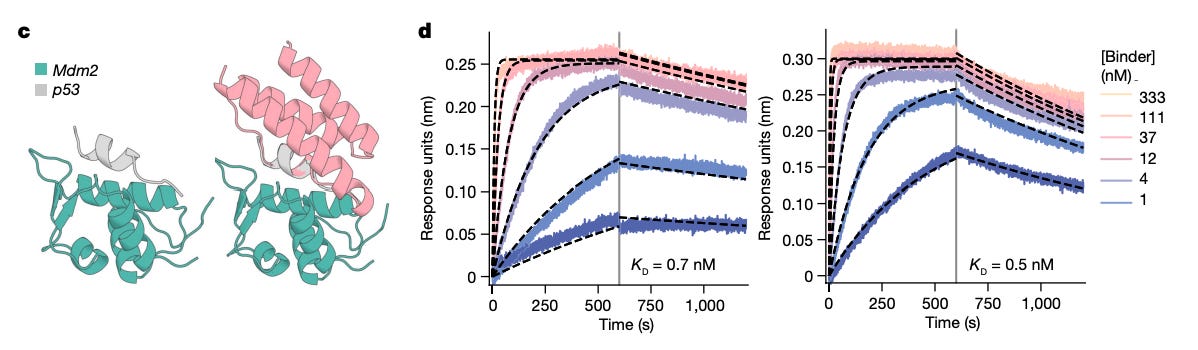
De novo binders, proteins designed from scratch to bind a specific target, are now routinely achieving nanomolar affinity on the first experimental round, without any optimisation. A 2024 study from Baker’s lab designed binders against a panel of therapeutic targets, including influenza and SARS-CoV-2, with success rates that would have been unthinkable five years ago.
Protein design competitions are providing independent validation. Adaptyv Bio, a cloud lab for protein designers based in Lausanne, ran an open EGFR binder competition in 2024 that benchmarked AI design methods head-to-head with standardised experimental testing. The results showed a 5x improvement in design success rates compared to earlier approaches, with some AI-designed binders outperforming clinical antibodies.
Stability is also improving. AI-designed proteins are increasingly more thermostable than their natural counterparts. They can be engineered to withstand higher temperatures and harsher conditions, which matters enormously for manufacturing and storage.
🧪 The Validation Bottleneck
Here’s the catch: designing a protein computationally takes hours. Testing it experimentally still takes weeks to months.
The field can now generate thousands of candidate designs per day. But each one needs to be synthesised, expressed, purified, and assayed to know if it actually works. That wet-lab step is the bottleneck, and it’s where a lot of promising computational designs die, not because the design was wrong, but because the testing pipeline can’t keep up.
This is driving a new category of infrastructure: automated, high-throughput protein testing platforms that can validate designs at the speed AI generates them. The goal is a closed loop, design, test, learn, redesign, running continuously with minimal manual intervention. We’ll be exploring this challenge in an upcoming deep dive with Adaptyv Bio, a cloud lab purpose-built for AI protein design validation. Stay tuned.
Until that loop is fully closed, the practical throughput of AI protein design is limited not by the models but by the experiments.
📊 Beyond Binders
Binding is the easiest thing to design for, because the objective is clear: does this protein stick to that target? But the field is pushing into harder problems.
Enzyme design, creating proteins that catalyse specific chemical reactions, is significantly more challenging because function depends on precise atomic arrangements in the active site, not just overall shape. Early results are promising but success rates are lower than for binders.
Multi-state design aims to create proteins that switch between conformations, molecular machines that respond to signals. This requires the model to optimise for multiple structures simultaneously, a much harder optimisation problem.
Symmetric assemblies, protein cages, rings, and lattices, are being designed for drug delivery, vaccine design, and materials science. RFDiffusion has demonstrated the ability to generate novel symmetric architectures that self-assemble when tested experimentally.
🔮 Where This Is Going
Three trends to watch.
Generative models are getting multimodal. The next generation of design tools will jointly generate structure and sequence, rather than treating them as separate steps. Models that can reason about structure, sequence, dynamics, and function simultaneously will produce better designs faster.
The data flywheel is spinning up. Every experimentally tested design, whether it works or not, generates training data that makes the next round of models better. Open repositories for protein design data are accelerating this. The more designs get tested, the faster the models improve.
The design-test loop is tightening. As automated testing platforms scale, the gap between computational design and experimental validation will shrink. The long-term vision is protein design on demand: specify the function you want, get a validated molecule back in days rather than months.
We’re still early. Most AI-designed proteins are relatively simple, single-domain binders tested in controlled settings. The gap between designing a protein that binds a target in a tube and one that works as a drug in a patient remains enormous. But the trajectory is clear: the tools are getting better, faster, and more accessible. And the proteins they’re producing are starting to work.
💬 Want to be featured in Kiin Bio Weekly?
Each issue we speak directly with researchers, scientists, and builders working at the frontier of AI in life sciences. If you're working on something in this space and think it would resonate with our community, I'd love to hear from you - fill out this form or reach out to me directly.
Found this useful? Forward it to a colleague in drug discovery or protein design - it's the best way to help the newsletter grow.
Subscribe now to stay at the forefront of AI in Life Science. Every week: primers, deep dives, and direct conversations with the people building the field.
Connect With Us
Have questions on this or suggestions for our next deep dive? We’d love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website