🧬 Adaptyv: Closing the Loop on Protein Design
Deep Dive | Edition 19
Welcome back to the deep dive, where we break down the AI tools and data reshaping how new drugs are discovered. In each edition, we speak directly with the teams behind these tools to explain what they solve, how they work and where they are going next.
Keeping up with AI x life science news can get exhausting.
It’s scattered across LinkedIn, X, Substack, arXiv, Slack, newsletters... and you still somehow miss the things that actually matter. Too much noise, not enough signal.
We’re building something to fix that: a smarter, more powerful way to stay on top of what’s actually relevant to you.
But we want to build it with you, not just for you. Take 2 minutes to tell us what’s missing. What you share will directly shape what we build, and you’ll be the first to benefit from it.
Today we’re looking at Adaptyv, a Lausanne-based startup that’s building the experimental infrastructure the protein design revolution has been missing. We spoke with Tudor, who leads community and protein engineering efforts at Adaptyv, about why designing proteins computationally was only ever half the problem, and what it takes to close the gap between prediction and proof.
“People are realising now that data is the bottleneck for what we can currently achieve in ML for protein design. They need specialised functional data and they need to generate it fast.”
— Tudor, Adaptyv
🔴 The Problem
AI protein design has exploded. Diffusion models, language models, and structure prediction tools can now generate novel protein sequences in minutes. But there’s a persistent bottleneck that sits downstream of all that computation: actually testing whether the designs work.
Before Adaptyv, a computational protein designer who wanted to validate their binders had limited options. You could work with a contract research organisation, with complex onboarding, long timelines, and months before you saw results. You could do it yourself if you were lucky enough to have lab access. Or you could be in one of a handful of major labs, like the Baker lab, that had the infrastructure to run binding affinity measurements at scale.
For everyone else, the growing wave of independent protein designers, small academic groups, and early-stage biotechs training their own generative models, experimental validation was a wall. You could design as many proteins as you wanted on a computer, but you had no efficient way to know which ones actually folded, bound their target, or did anything useful.
“It was quite a black and white situation,” Tudor explains. “You were either in one of the bigger labs or you weren’t. And if you weren’t, you were designing stuff on the computer with no idea what it does in the real world.”
💡 The Idea
Adaptyv’s answer: a cloud lab purpose-built for protein designers.
The thesis has been core to the company since its founding by Julian Englert and Daniel Nakhaee-Zadeh, both engineers from EPFL. They initially built a microfluidics platform for high-throughput antibody-antigen binding testing, but when that approach proved too experimental, they pivoted. First to BLI and SPR-based measurement approaches in late 2023, then to an official platform launch in 2024.
The model is simple. You sign up on the Foundry platform. Upload your sequences via CSV. Add details: tags, antibody formats, terminal modifications. Select your target from a catalogue sourced from multiple suppliers. Submit. Adaptyv handles everything else.
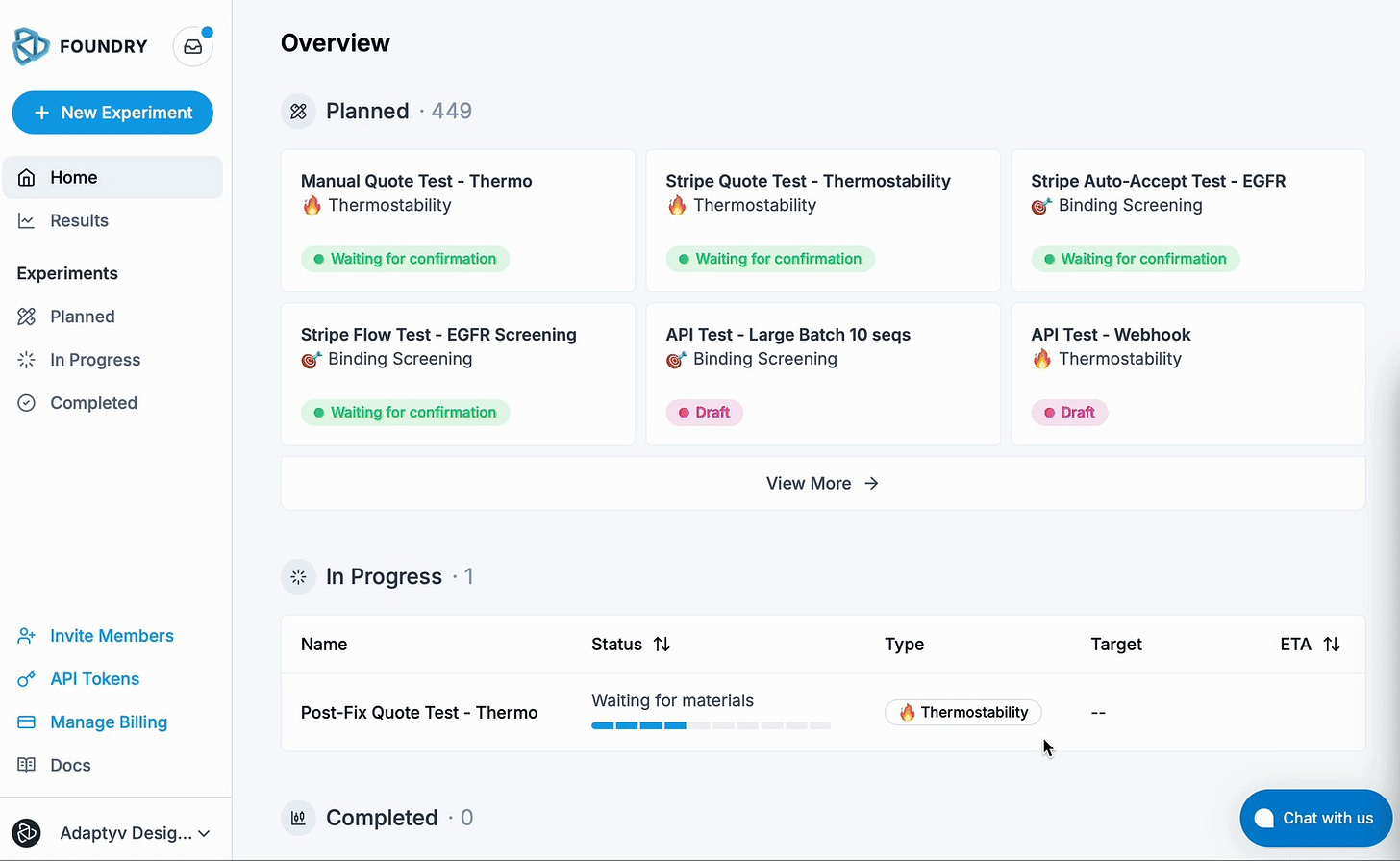
The Foundry experiment creation workflow: choose your desired assay, choose targets for binding experiments, number of replicates, upload your sequences, and get an experiment quote for your draft experiment.
Behind the interface sits what the team calls LabOS, an orchestration brain that manages scheduling, expression, measurement, curve fitting, quality control, and results delivery across their automated systems. Users get back visualisations, raw sensorgrams, fitted binding curves, and all underlying data in a single package.
The key technical innovation is miniaturisation. By running cell-free expression reactions in microlitre volumes rather than large batch cultures, Adaptyv achieves roughly a 1,000x reduction in reagent use. No recombinant E. coli. No multi-day protein expression waits. Just small, fast, cell-free reactions that produce enough protein to measure on BLI or SPR.
📊 The Data
Current turnaround is approximately two weeks: one week for target QC if the target is new, then one week for the experiments themselves. The goal is to push that down to two to three days, and ultimately to near-instant validation that matches the speed of ML training cycles.
“Ideally, you want to get the experimental validation within the same time pressure as a training run,” Tudor says. “Almost instant. If we could make it faster, we would.”
The platform isn’t just a validation service; it’s becoming a data engine. Some customers are already running active learning loops: generate a set of designs, send them to Adaptyv, get results, retrain, repeat. Others are running large-scale campaigns to map the druggability of entire target spaces.
A typical use case: a team trains a diffusion model on PDB data, generates novel binders, and validates them through Adaptyv. The experimental results feed back into the next round of model training. Each cycle produces better designs and richer data.
“Both work hand in hand,” Tudor explains. “Now people are using us primarily for validation. But ultimately we want to go in the direction of large-scale custom data generation campaigns. We want to be a protein data centre.”
🔬 Proteinbase
The data Adaptyv was generating led naturally to their second platform: Proteinbase.
The problem it addresses is fragmentation. Existing protein databases use different protocols, different measurement methods, and different standards. Data from one source is hard to compare with data from another. Teams trying to train models on aggregated functional protein data spend enormous effort just standardising datasets, and even then, results often don’t reproduce.
“People were saying that what they’re missing is a unified database for functional proteins with standardised protocols,” Tudor says. “Everyone was trying to standardise datasets from different sources and running into the same problems. At least if everything is from a single source, it’s better for training better models.”
Proteinbase hosts competition data, provides unified downloads, and is building out community features: leaderboards, badges, knowledge sharing. It’s part database, part competition platform, part community hub.
The competitions have been a major growth driver. Recent rounds attracted over 600 participants, with hit rates up to 13% and binders in the nanomolar range. Adaptyv has also been hosting hackathons internationally in San Francisco and Berlin, with more planned across Europe. A recent GEM Bio Workshop competition hosted at ICLR on a disordered target (RBX1) drew more than 180 submissions.
💊 Who It’s For
Adaptyv serves three overlapping audiences: academic groups training and validating protein design models, biotech and pharma teams running lead optimisation or target characterisation campaigns, and independent protein designers who previously had no access to experimental validation.
Pricing is transparent and visible on the platform.
The philosophy is democratisation. Protein design has historically been concentrated in a small number of elite labs. Adaptyv is opening that up, not just through the lab infrastructure but through the community.
“Protein design has been super insular,” Tudor says. “It was either you were in one of the bigger labs or you weren’t. Giving people access to experimental validation and a community to share what they’ve learned, that’s how you grow the field.”
🔮 The Future
The next 12 months are focused on speed and automation.
On the Adaptyv platform: two-day experimental results, an expanded assay array, more standardisation, and deeper investment in agent-based workflows. The API is already designed for programmatic experiment submission, and the team is building towards a future where AI agents can control every aspect of the pipeline, from sample handling to liquid handlers to results retrieval.
“We have a two-level system,” Tudor explains. “A higher-level customer API for submitting experiments and getting results, and a lower-level API for every single machine interaction. In the future, agents could control all of it.”
On Proteinbase: more visible content, international hackathon expansion across Europe, and continued growth of the competition platform. The demand is clear: organisers are reaching out to Adaptyv to add protein design tracks to their events, and recent competitions have been oversubscribed.
The longer-term vision ties both platforms together. As the field recognises that data, not compute or model architecture, is the binding constraint on progress in ML for protein design, Adaptyv is positioning itself as the infrastructure layer that generates, validates, and distributes that data at scale.
Design is only half the problem. Adaptyv is building the other half.
👨🔬 Get in touch with Tudor.
Thanks for reading Kiin Bio Weekly!
💬 Get involved
We’re always looking to grow our community. If you’d like to get involved, contribute ideas or share something you’re building, fill out this form or reach out to me directly.
Subscribe now to stay at the forefront of AI in Life Science and keep up with this upcoming season of deep dives.
Connect With Us
Have questions on this or suggestions for our next deep dive? We’d love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website