🧬 A Primer on AI Agents in Drug Discovery
The data exists. The tools exist. The problem is getting them to talk to each other.
Welcome back to your weekly dose of AI news for Life Science!
What’s your biggest time sink in early drug discovery process?
Drug discovery has a productivity problem, and it is not the one people usually talk about.
The bottleneck is rarely a shortage of data or computational tools. It is the gap between them. You have a model that predicts toxicity, another that generates novel structures, a third that searches the literature. None of them talk to each other. Every handoff between tools is a handoff between people, and people are slow.
We spoke to Srijit Seal, visiting scientist at the Broad Institute of MIT and Harvard and lead author of a new paper on agentic AI in drug discovery, about what these systems can actually do today, where pharma is in adopting them, and why he thinks the next decade looks very different from the last.
“As LLMs get better at reasoning, can they determine which tools should be used for what kind of tasks? That’s the core question we set out to explore.”
— Srijit Seal
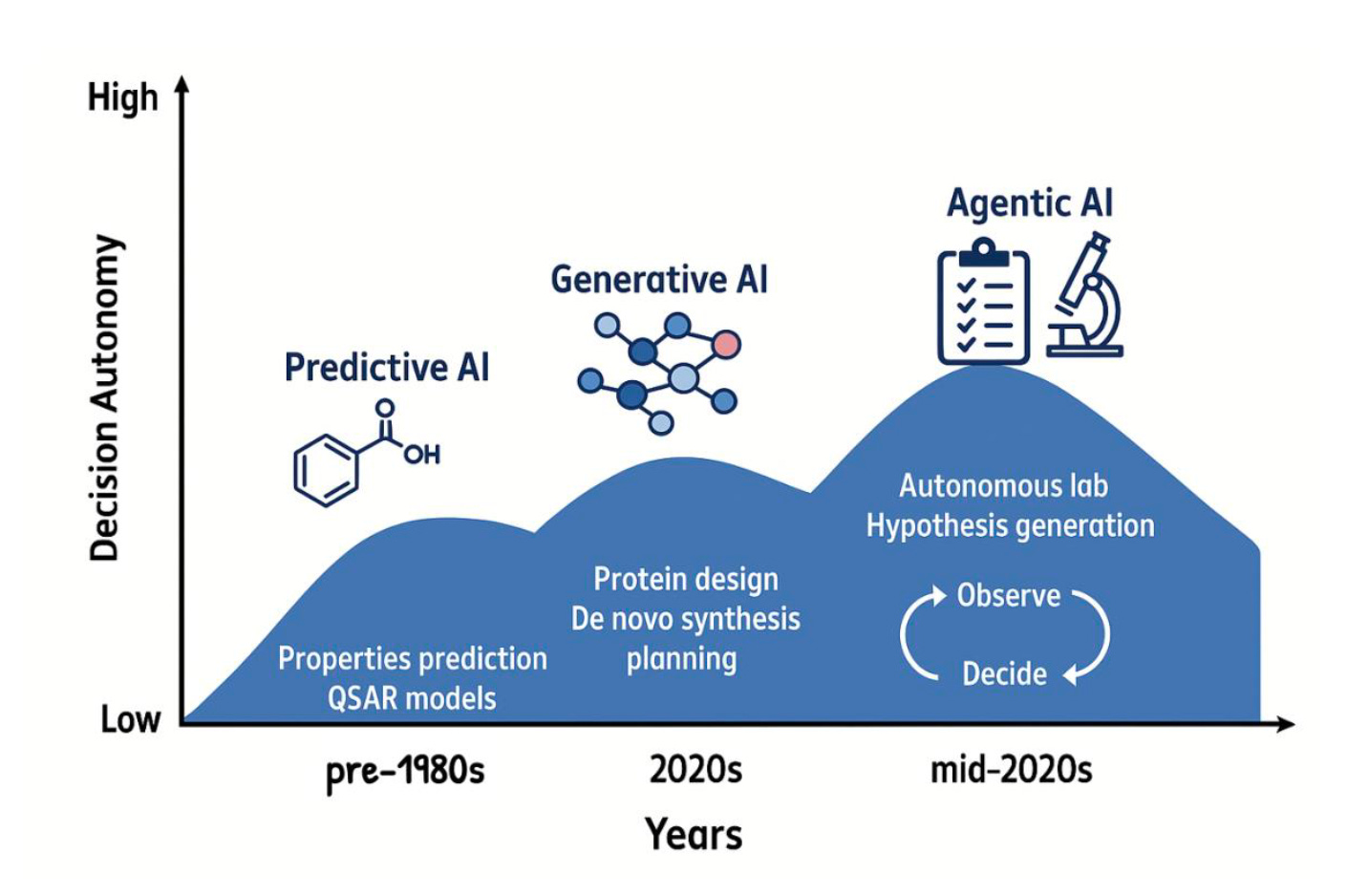
🤖 What Makes AI “Agentic”?
Before getting into applications, it is worth pinning down what the word means, because it gets thrown around loosely. Seal’s working definition is practical: an agentic system has access to tools and makes decisions about which ones to use. It executes functions. It does not just produce text.
“They have to have access to tools and make decisions on which tools to use,” he explains. “Not just summarise the next token and write down text. We need them to constantly use some tools to inform what they want to write next.”
That distinction matters quite a lot. A language model that summarises a paper is useful. An agent that retrieves the paper, extracts the relevant assay data, cross-references it against a compound database, runs a toxicity prediction, and writes the summary is a different category of thing. The first is a tool. The second is closer to a junior scientist.
Seal and his co-authors identify four tool types that matter in drug discovery: perception (querying databases, retrieving literature), computation (running predictions, simulations, docking), action (controlling lab hardware), and memory (storing findings across sessions so the agent builds on what it has already learned). Together they enable a closed loop: gather evidence, compute predictions, run experiments, remember results, and repeat.
🏗️ The Architectures Worth Knowing
How you wire agents together shapes what they can do. The simplest design is ReAct, short for Reasoning and Acting: the model reasons, calls a tool, observes the result, reasons again. Fast and flexible, good for focused tasks like literature triage or compound queries. Its weakness is coherence on tasks that sprawl across many steps.
Reflection architectures run multiple models in dialogue, each critiquing the other. Slower and more compute-intensive, but meaningfully better for tasks that need strategic rigour, like mapping out a multi-step synthetic route. Supervisor architectures model a research team: one orchestrating agent breaks a complex task into pieces and delegates to specialist sub-agents, each with its own domain. Swarm architectures remove the central coordinator entirely, so agents communicate peer-to-peer. That scales well across institutions, though coordination overhead grows with it.
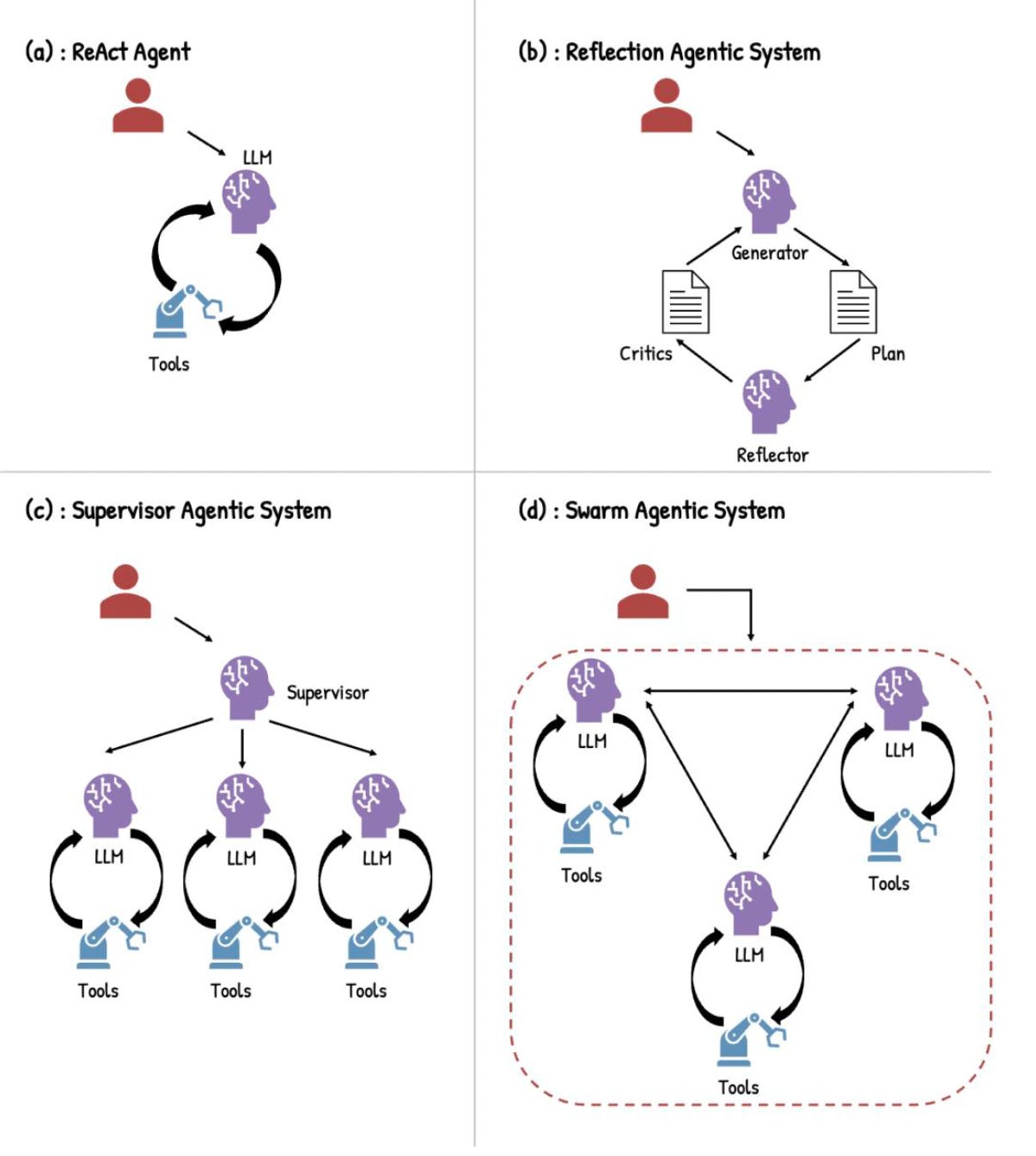
For most of the real-world applications Seal describes, supervisor architectures do the heavy lifting. Drug discovery spans biology, chemistry, clinical data, and business context. No single agent holds all of it.
⚙️ What Agentic AI Is Already Doing
Seal’s paper presents case studies from seven startups, including ourselves, working across the drug discovery pipeline.
In literature and structure-activity relationship analysis, a supervisor-based system compressed weeks of manual review into hours, retrieving kinase selectivity data, identifying assay inconsistencies, and producing structured, cited reports.
In toxicity assessment, a ReAct-based agent combined predictive toxicology models with literature retrieval to evaluate endocrine disruption risk, reconciling computational findings with regulatory evidence in a fraction of the usual time.
In lab automation, an agent designed an automated qPCR assay for AAV quantification. Within two hours it generated a curated literature review, a MIQE-aligned protocol, and executable robot code, a process that typically takes one to four months.
More ambitiously, integrated platforms are beginning to run multi-step preclinical workflows: disease landscape analysis, target prioritisation from public omics data, structural modelling, and ranked hit generation, all in hours rather than weeks.
The common thread is not intelligence in isolation, but orchestration across tools.
⚠️ Why This Is Still Hard
None of this means the problem is solved. Seal is direct about where the friction is.
Drug discovery data is unusually difficult to reason over. Unlike images or text, it is deeply conditional: the measured activity of a compound depends on the assay format, the concentration, the cell line, the lab running the experiment. The same molecule can produce IC50 values that differ by an order of magnitude across the literature, not because the measurements are wrong, but because they are measuring slightly different things under different conditions. Getting an agent to reason reliably across this kind of data requires careful engineering, not just a capable model.
Security is a genuine concern. Agentic systems that browse external sources are vulnerable to prompt injection: malicious instructions embedded in a document or webpage that cause the agent to take unintended actions. When your agent has access to proprietary compound libraries and can exfiltrate data externally, the attack surface is not theoretical. Hallucinations, while less frequent in modern models, remain a risk in high-complexity scenarios, exactly the kind where agentic AI is most useful. And there is no mature benchmarking framework for agentic drug discovery systems yet, which makes it hard to compare capability claims rigorously.
💊 Where Pharma Is
The gap between what these systems can demonstrate and what is running inside large pharmaceutical companies is wide. That is not a criticism; it reflects appropriate caution in an industry where errors have consequences for patients.
Seal sees a clear pattern: initial scepticism from scientists and programmers, rapid conversion once they try the tools on real data, slow movement through institutional sign-off. “Once they try it on dummy data, they are like: this is actually quite good. I can automate a lot of my work.” The bottleneck is not conviction. It is process.
Traction right now is concentrated in lower-stakes tasks: literature summarisation, report drafting, document search. One underappreciated effect is democratisation: agentic tools give a small biotech with one computational scientist access to capabilities that previously required a whole team. “Previously you would have one person telling me to dock on AlphaFold, another to predict toxicity, another to automate the robot,” Seal says. “It is quite difficult to find someone with all those skills.”
The companies investing most visibly, AstraZeneca, Eli Lilly, GSK, are largely building internal infrastructure and running pilots. Industry watches academic and startup work carefully and pulls selectively. “Some companies will wait to see which one Lilly chose. If a bigger company signs a deal, the others follow.”
🔮What Comes Next
The near-term direction is self-driving laboratories: closed-loop systems that design experiments, instruct hardware, analyse results, and update hypotheses without human sign-off at each step. Prototypes exist. Scaling them to multi-step workflows with the reliability pharma demands is the engineering challenge.
New infrastructure is arriving to support agent collaboration across organisations. Anthropic’s Model Context Protocol (MCP) and Google’s Agent2Agent (A2A) protocol, now consolidated under the Linux Foundation, are building standardised interfaces so agents from different companies and labs can exchange data regardless of their underlying stack. The longer-term vision is discovery pipelines where a target identification agent at one institution hands off to a synthesis agent at another without a human in the middle.
Seal’s most striking prediction is what he calls the agentic portfolio. A scientist’s accumulated knowledge, preferred protocols, learned intuitions about which assay conditions are reliable, encoded as instructions that travel with them between jobs. When you hire someone, you hire their agents too. It sounds futuristic until you consider that this already happens informally whenever a scientist moves labs and brings their scripts, contacts, and institutional memory. The agents just make it explicit.
The consistent theme across all of it: these tools amplify human judgement, they do not replace it. The scientist who knows why a synthesis route is problematic, who recognises when a model’s training data does not reflect the patient population, who understands the difference between a statistically significant and a clinically meaningful result: that expertise remains the rate-limiting step. Agentic AI handles the orchestration. The thinking that matters most is still ours.
Thanks for reading Kiin Bio Weekly!
💬 Get involved
We’re always looking to grow our community. If you’d like to get involved, contribute ideas or share something you’re building, fill out this form or reach out to me directly.
Subscribe now to stay at the forefront of AI in Life Science and keep up with this upcoming season of deep dives.
Connect With Us
Have questions on this or suggestions for our next deep dive? We’d love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website

