Arc Institute's MULTI-evolve, EleutherAI's Deep Ignorance, and UPenn/Stanford's iSight
Kiin Bio's Weekly Insights
Welcome back to your weekly dose of AI news for Life Science!
What’s your biggest time sink in the drug discovery process?
MULTI-evolve: Accelerating protein engineering through predictive epistasis
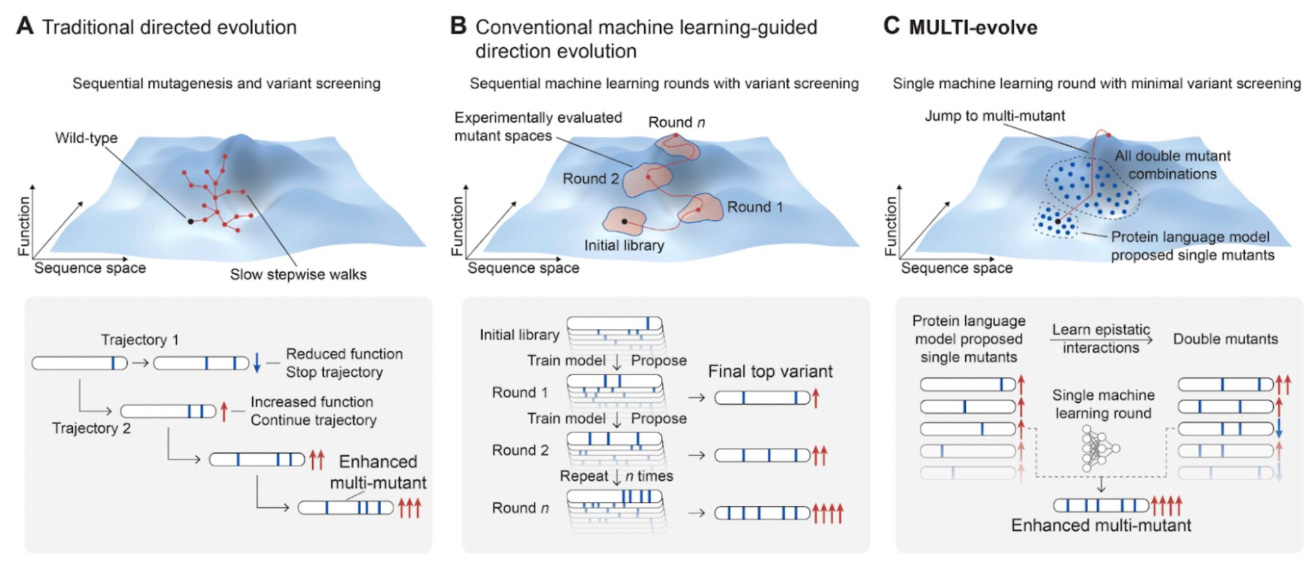
🔬 Protein engineering traditionally requires multiple rounds of directed evolution to identify beneficial mutations, a time-consuming and resource-intensive process.
Arc Institute’s MULTI-evolve bridges predictive biological modelling with functional laboratory optimisation to accelerate protein engineering. By integrating computational mutation prediction with targeted experimental validation, the framework identifies and combines beneficial mutations without requiring repeated rounds of directed evolution.
🧬 MULTI-evolve addresses epistasis - where mutation effects depend on one another - by experimentally testing pairwise mutation combinations and training models to predict higher-order interactions. This enables efficient design of synergistic multi-mutant variants with substantially enhanced activity.
⚡ The framework was applied to APEX, CRISPR-Cas13d, and an anti-CD122 antibody. MULTI-evolve first identified improved single mutants (up to threefold improvement) and leveraged epistatic modelling to design variants containing up to seven mutations, achieving improvements of up to 256-fold.
Applications & Insights
1️⃣ Improving Discovery of Function-Enhancing Mutations Combining protein language models with targeted validation, MULTI-evolve prioritises beneficial single mutations before combinatorial optimisation begins.
2️⃣ Modelling of Epistasis Quantifying epistatic interactions via pairwise mutation testing enables prediction of synergistic multi-mutant variants without extensive screening.
3️⃣ Single Round Design of High-Order Mutants Single-cycle generation of multi-mutant variants accelerates development, reduces laboratory workload, and eliminates large combinatorial libraries.
4️⃣ Multi-Mutant Engineering of Diverse Proteins MULTI-evolve has enhanced APEX via a valine substitution improving hydrophobicity and sodium ion coordination, engineered a dCas-Rx splicing tool with improved RNA binding and crRNA recognition, and demonstrated promise in multi-objective antibody design with improved binding and expression.
💡 Why This Is Cool
MULTI-evolve unites predictive modelling with targeted experimental validation to streamline protein engineering. By quantifying epistatic interactions, it enables single-cycle optimisation, reduces laboratory burden, and demonstrates broad utility across enzymes, CRISPR tools, and antibodies.
📖 Read the paper
💻 Try the code
Deep Ignorance: Filtering dangerous knowledge at training time
🔒 What if the safest AI model is simply one that never learned the dangerous stuff in the first place? That’s the question researchers at EleutherAI, the UK AI Security Institute, and Oxford’s OATML group decided to take seriously and honestly, the results are pretty compelling.
Here’s the thing: open-weight models can be downloaded and fine-tuned by anyone. Every post-training safety fix tried so far gets undone within a few hundred steps. The weights remember what they saw during training, and that memory is surprisingly easy to recover.
🧪 Applications and Insights
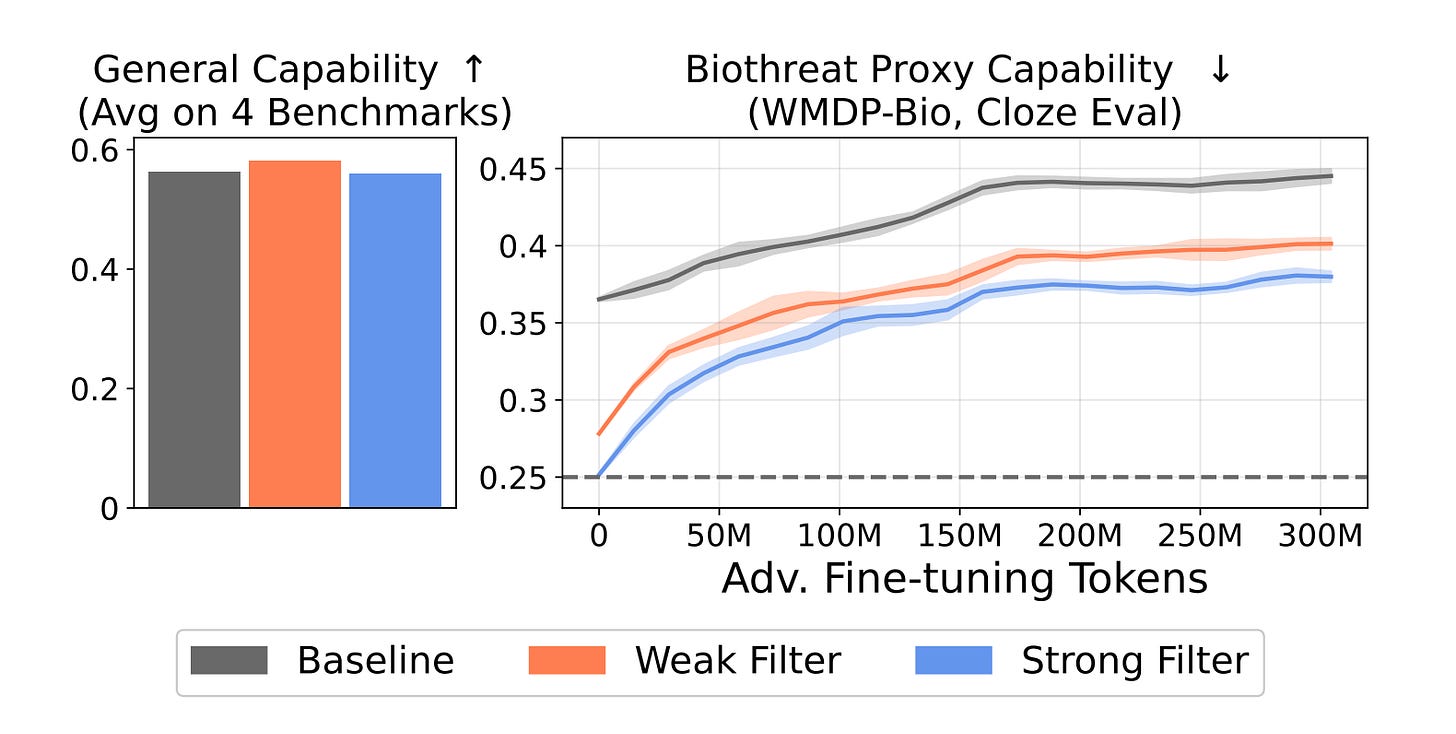
1️⃣ Tamper-resistant biothreat suppression Filtered 6.9B models held up against 10,000 fine-tuning steps and 300M tokens of adversarial biothreat text, more than ten times the resistance of any post-training baseline ever tested.
2️⃣ No capability trade-off Filtering costs less than 1% of total training FLOPs. Performance on MMLU, HellaSwag, PIQA, and LAMBADA? Basically flat.
3️⃣ Filtering and Circuit Breaking are better together Filtering alone can’t stop in-context retrieval attacks. Pair it with Circuit Breaking and that gap closes. Neither works alone, but together they’re genuinely robust.
4️⃣ It doesn’t work for everything Filtering behavioural propensities like toxicity and jailbreak compliance is a different beast. Filtered models can actually get more vulnerable to few-shot attacks on those tasks. Knowledge and behaviour are not the same problem.
💡 Why This Is Cool
The field has spent years trying to unlearn dangerous knowledge after the fact. This paper just asks: why teach it at all? Simple idea, rigorous execution, and refreshingly honest about where it falls short.
📖 Read the paper
💻 Try the code
iSight: Automated IHC staining assessment at scale
🔬 Stanford Healthcare runs over 3,500 immunohistochemistry cases every single month. Back in 2005, that number was 700. The workload has quintupled and the workforce hasn’t kept up. You know what that means in practice: more pressure, more variability, more room for error.
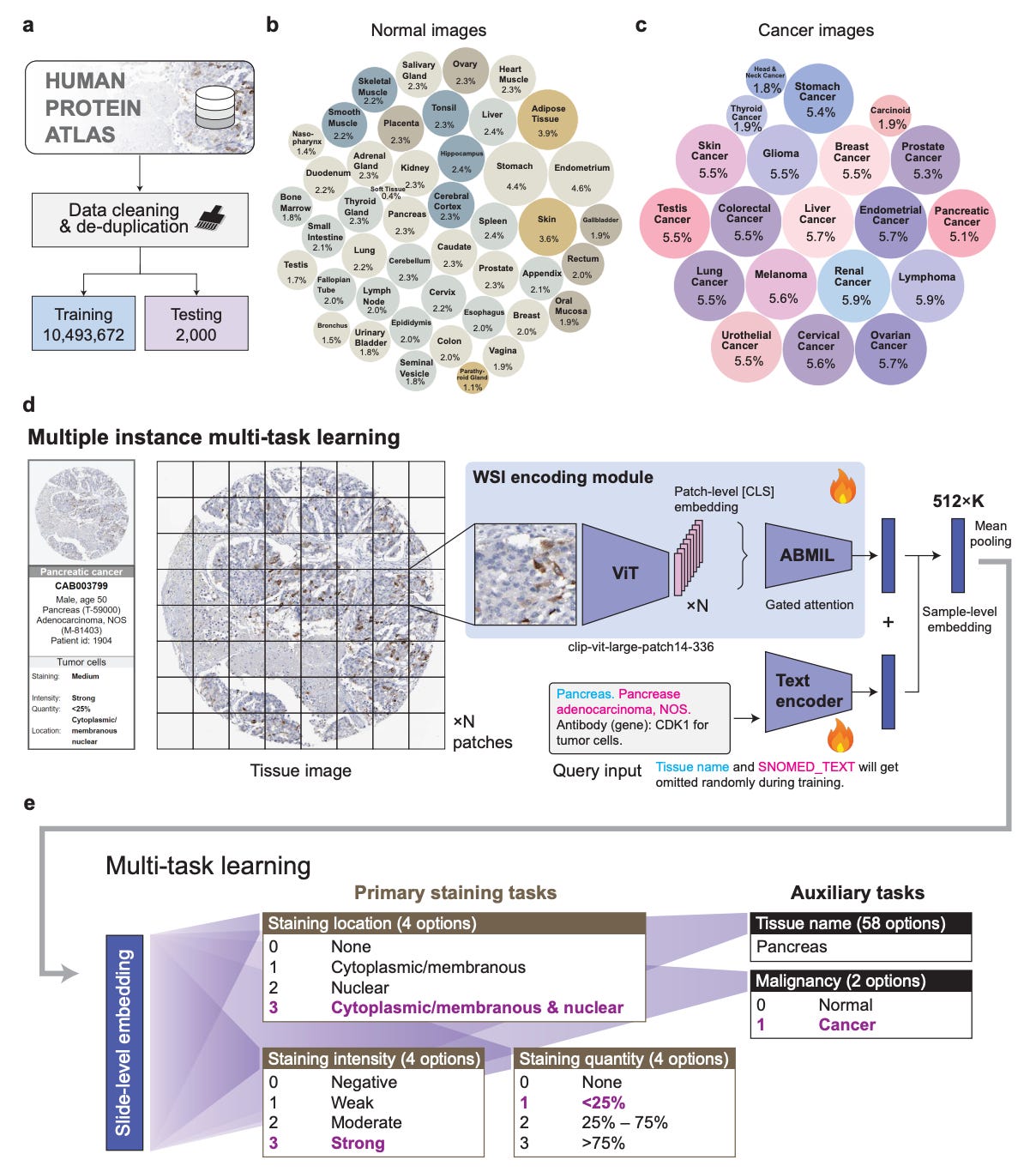
Researchers at the University of Pennsylvania and Stanford built iSight to help. It’s a multi-task AI model for automated IHC staining assessment, trained on HPA10M, a freshly curated dataset of over 10 million IHC images across 45 normal tissue types and 20 cancer types. Nothing like it has existed at this scale before.
🧫 Applications and Insights
1️⃣ Three staining tasks, one model iSight predicts intensity, localisation, and quantity simultaneously, hitting 85.5% accuracy for location, 76.6% for intensity, and 75.7% for quantity, beating fine-tuned PLIP and CONCH by 2.5–10.2%.
2️⃣ Calibration you can actually trust Expected calibration errors sat between 0.015 and 0.041. The confidence scores reflect real accuracy, which matters enormously when a clinician is deciding whether to act on a prediction.
3️⃣ It makes pathologists more consistent Eight pathologists evaluated 200 images before and after seeing iSight’s suggestions. Inter-rater agreement jumped from Cohen’s κ 0.63 to 0.70. The AI didn’t override anyone, it just quietly anchored the room.
4️⃣ Holds up under messy real-world conditions Salt-and-pepper noise, tissue fold artefacts, scan dropouts, tested across four severity levels, performance never deviated more than 1.5% from baseline.
💡 Why This Is Cool
Most pathology AI papers benchmark and stop there. This one actually put the model in front of clinicians and watched what happened. The finding that pathologists improve but still trail the AI alone is genuinely fascinating, it says as much about human decision-making as it does about the model.
📖 Read the paper
💻 Try the code
🗓️ Events & Competitions
We’re launching a new section of the newsletter dedicated to tracking the best competitions, hackathons, and community challenges happening!
If you’re looking to get hands-on with cutting-edge targets, and have your work experimentally validated, these are the events worth your time. We’ll be featuring these regularly, so if you know of something worth highlighting, reply and let us know!
First up, two hackathons happening this weekend courtesy of our friends at Adaptyv Bio 👇
bioArena x Adaptyv: Agents for Protein Design Hackathon | Feb 28, San Francisco
Teams are using AI agents to design binders against TREM2, one of the strongest genetic risk factors for Alzheimer’s disease and a highly active area of drug discovery. Top designs get experimentally validated by Adaptyv in the wet lab.
Berlin Bio x AI Hackathon | Feb 27-28, Berlin
24 hours, an aging-relevant target, and $10,000 in Adaptyv lab credits on the line. The top 100 designs go straight to Adaptyv’s wet lab for experimental validation, with additional tracks on genome modeling and agentic AI in life sciences. Applications are closed but we’ll be watching the results closely.
Thanks for reading!
💬 Get involved
We’re always looking to grow our community. If you’d like to get involved, contribute ideas or share something you’re building, fill out this form or reach out to me directly.
Connect With Us
Have questions or suggestions? We'd love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website