Loschmidt Labs' TmProt, UVA's YuelDesign, and Caltech's DISCO
Kiin Bio's Weekly Insights
Welcome back to your weekly dose of AI news for Life Science!
Keeping up with AI x life science news can get exhausting.
It’s scattered across LinkedIn, X, Substack, arXiv, Slack, newsletters... and you still somehow miss the things that actually matter. Too much noise, not enough signal.
We’re building something to fix that: a smarter, more powerful way to stay on top of what’s actually relevant to you.
But we want to build it with you, not just for you. Take 2 minutes to tell us what’s missing. What you share will directly shape what we build, and you’ll be the first to benefit from it.
🇺🇸 Meet Us at Bio-IT, Boston
🤝 Our CEO Filippo and CTO Bogdan will be at Bio-IT World in Boston. If you are attending and would love to connect in person, reach out and let us know.
TmProt 1.0: Predicting Protein Melting Temperatures for Enzyme Discovery
🔬 Predicting protein melting temperatures (Tm) is critical for enzyme engineering: thermostable enzymes last longer and tolerate harsher conditions. But most AI predictors train on mass-spectrometry data from whole cells, which is fundamentally different from purified protein measurements. The Loschmidt Labs team found near-zero correlation (r = 0.05) between proteomics and biophysical Tm datasets for the same proteins.
TmProt 1.0 from Loschmidt Laboratories (Masaryk University) fixes this by training exclusively on biophysically measured melting temperatures. Built on ESM-2 fine-tuned with LoRA, it is now live on Hugging Face.
🧬 The team assembled ProMelt, a curated set of 45,441 proteins with experimental Tm values, validated across 5 independent biophysics test sets. TmProt outperforms TemBERTure, DeepSTABp, and SaProt across most benchmarks.
⚡ Absolute Tm regression remains hard, but TmProt excels where it matters most: ranking thermostable candidates. For classifying proteins with Tm at or above 60 degrees C, it achieves an AUC of 0.75, and is fully integrated into EnzymeMiner 2.0 for end-to-end enzyme mining.
🔬 Applications and Insights
1️⃣ Thermostable Enzyme Enrichment
Best used as a ranking tool to prioritise thermostable candidates from large sequence sets, making it practical for early-stage enzyme discovery campaigns.
2️⃣ Training Data Matters More Than Architecture
The near-zero correlation between proteomics and biophysical Tm explains why previous predictors underperformed. Curating ProMelt was the single biggest driver of improvement.
3️⃣ Lightweight and Accessible
LoRA fine-tuning keeps the model efficient enough to run as a free web server, lowering the barrier for labs without GPU infrastructure.
4️⃣ Integrated Discovery Pipeline
Full integration with EnzymeMiner 2.0 means researchers go from sequence database to stability-ranked shortlist in one workflow.
💡 Why This Is Cool
Data quality beating model complexity. The proteomics vs biophysical Tm disconnect explains years of underwhelming predictions. By fixing the data rather than stacking layers, TmProt delivers where enzyme engineers actually need it.
🌐 Try TmProt.
🌐 Try EnzymeMiner 2.0.
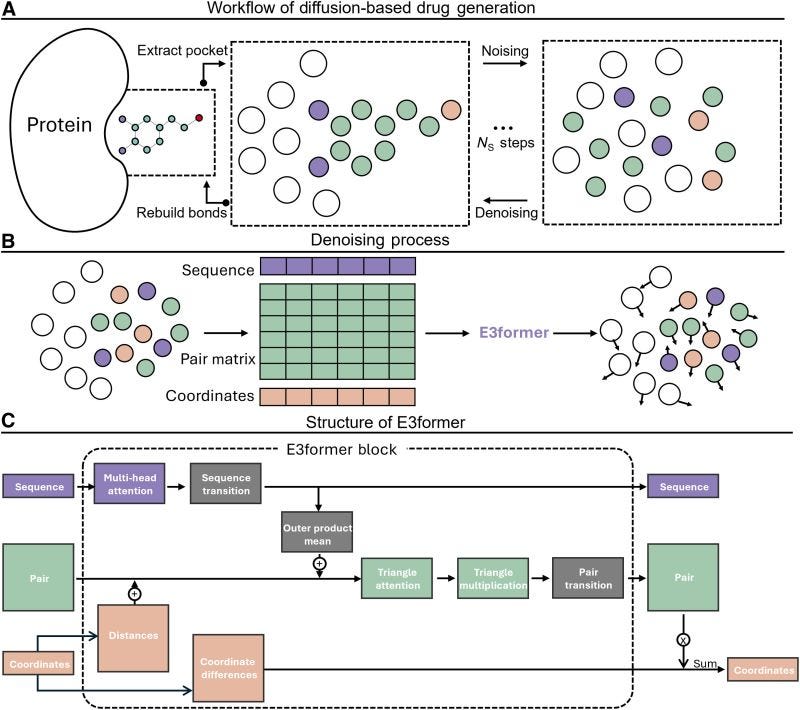
YuelDesign: Diffusion-Based Molecule Design in Flexible Protein Pockets
🔬 Proteins undergo conformational changes upon ligand binding, yet most deep learning generative models treat pockets as rigid, generating molecules for a single frozen conformation. The pocket shape in a crystal structure may not be the one your molecule actually binds.
YuelDesign from the Dokholyan lab at the University of Virginia jointly models both the pocket structure and ligand conformation, allowing protein and molecule to co-adapt during generation.
🧬 Two diffusion processes run simultaneously: an elucidated diffusion model (EDM) for 3D coordinates and a discrete denoising diffusion model (D3PM) for atom types. Both use E3former to maintain rotational and translational equivariance.
⚡ The result is molecules with favourable drug-likeness, low synthetic complexity, diverse functional groups, and docking energies comparable to native ligands. By letting the pocket breathe during generation, YuelDesign captures induced-fit effects that rigid methods miss.
🔬 Applications and Insights
1️⃣ Induced-Fit Drug Design
Jointly diffusing pocket and ligand explores conformational states that only emerge upon binding, capturing selectivity-driving dynamics rigid models cannot access.
2️⃣ Synthesisable and Drug-Like Output
Generated molecules score well on synthetic accessibility out of the box, reducing the gap between computational hits and what a medicinal chemist would actually make.
3️⃣ Diverse Chemical Exploration
D3PM atom-type diffusion produces varied functional groups rather than collapsing to a narrow series, giving broader chemical space coverage per run.
4️⃣ Beyond Rigid Docking
Treating flexibility explicitly moves generative design closer to real protein-ligand recognition, where both partners adjust shape upon binding.
💡 Why This Is Cool
The rigid-pocket assumption has limited structure-based design for decades. YuelDesign tackles it with a clean dual-diffusion architecture that still produces drug-like, synthetically accessible molecules. A meaningful step for generative molecular design.
📄 Read the paper.
💻 Try the code.
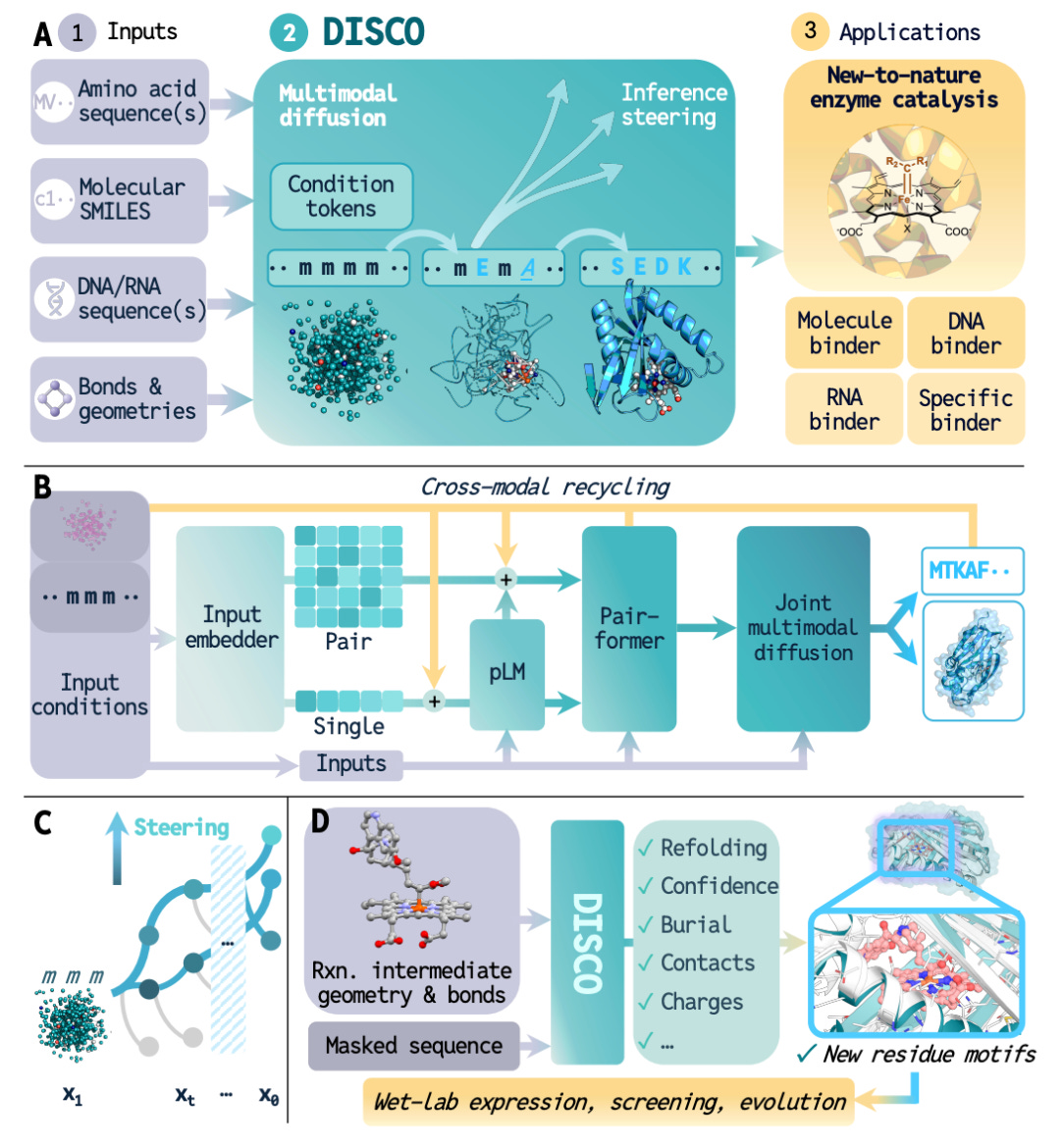
DISCO: AI-Driven Co-Design of Enzyme Sequence and Structure for Drug Discovery
🔬 Traditional enzyme design starts with a theozyme: a hand-crafted catalytic geometry based on detailed mechanistic knowledge. This works when you understand the mechanism, but limits you to chemistries where that knowledge exists.
DISCO (Diffusion for Sequence-Structure Co-design) from Caltech co-generates sequence and structure simultaneously using diffusion over discrete amino acids and continuous 3D coordinates, conditioned only on DFT-derived reactive intermediates. No predefined catalytic motifs needed.
🧬 Given a target chemistry, DISCO explores catalytic solutions freely, generating novel active sites and repurposing unrelated protein folds for catalysis. The model jointly optimises sequence and structure in a single pass, preserving their natural interdependence.
⚡ On the STUDIO-179 benchmark, DISCO generated the highest proportion of co-designable protein-ligand complexes for 178 of 179 cases. High-performing enzymes were found from just 90 designs, and for C(sp3)-H insertion, a poorly understood reaction, DISCO-designed enzymes matched variants from extensive directed evolution.
🔬 Applications and Insights
1️⃣ Bypassing Theozyme Design
Conditioning on reactive intermediates rather than predefined geometries lets DISCO tackle chemistries where mechanistic knowledge is incomplete or unavailable.
2️⃣ Evolvable by Design
Generated enzymes respond well to directed evolution, meaning computational design and experimental optimisation work as complementary stages.
3️⃣ Novel Folds for Novel Chemistry
DISCO repurposes unrelated protein scaffolds and generates new active sites, expanding the design space beyond known enzyme families.
4️⃣ Reduced Experimental Screening
Functional enzymes from just 90 designs dramatically reduces screening burden compared to traditional library approaches.
💡 Why This Is Cool
Enzyme design has always been bottlenecked by mechanistic knowledge. DISCO removes that constraint. Matching directed evolution on a poorly understood reaction from de novo designs suggests AI can now explore catalytic solutions humans would not think to try.
📄 Read the paper.
💻 Try the code.
🗓️ Events & Competitions
The best competitions, hackathons, and community challenges in AI x life sciences, curated weekly. Know something worth featuring? Reply and let us know.
More upcoming events:
Agentic Genomics: Hands-on with AI for Variant Interpretation and GWAS | April 22, Virtual
A free, two-session workshop co-hosted by Manuel Corpas and Segun Fatumo, running entirely in Google Colab. No setup, no cost, just a browser. Session one covers variant interpretation using Ensembl VEP, ClinVar, and ACMG criteria. Session two runs a full GWAS pipeline with ClawBio, including polygenic risk scores and locus fine-mapping. Built to remove barriers for researchers anywhere in the world.
Register (free): https://luma.com/vc98zeik
BioHackathon Europe 2026 | November 9-13, Barcelona
ELIXIR’s annual international bioinformatics hackathon, running since 2018. 160+ participants, five days of collaborative coding on open bioinformatics infrastructure and tools. The call for project proposals opens March 16 and closes April 15 - so if you want to lead a project, that’s your window.
Thanks for reading!
💬 Get involved
We’re always looking to grow our community. If you’d like to get involved, contribute ideas or share something you’re building, fill out this form or reach out to me directly.
Connect With Us
Have questions or suggestions? We'd love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website