Stanford's CellVoyager, Oxford's MacroGuide, and Arc Institute's BioReason-Pro
Kiin Bio's Weekly Insights
Welcome back to your weekly dose of AI news for Life Science!
Keeping up with AI x life science news can get exhausting.
It’s scattered across LinkedIn, X, Substack, arXiv, Slack, newsletters... and you still somehow miss the things that actually matter. Too much noise, not enough signal.
We’re building something to fix that: a smarter, more powerful way to stay on top of what’s actually relevant to you.
But we want to build it with you, not just for you. Take 2 minutes to tell us what’s missing. What you share will directly shape what we build, and you’ll be the first to benefit from it.
CellVoyager: An AI agent for autonomous biological data analysis
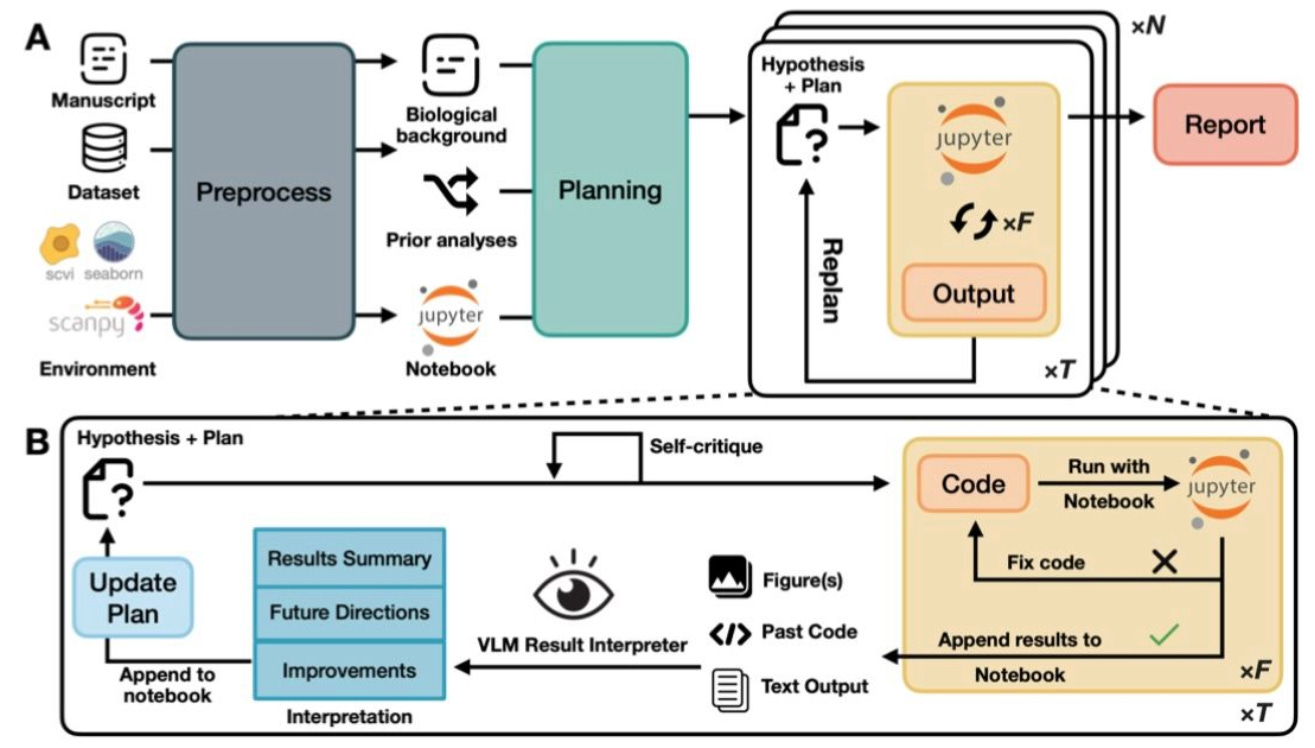
🔬 Most AI tools for single-cell analysis execute the analyses a user asks for. CellVoyager does the opposite: given a processed scRNA-seq dataset and prior analyses from a published paper, it autonomously proposes and executes novel analytical directions that build on existing work.
Stanford University’s Zou Lab built CellVoyager on LLMs running within a fixed Jupyter environment. It generates iterative “exploration blueprints” - self-critiqued analytical plans executed step-by-step with automatic code fixing and VLM-based result interpretation.
🧬 CellVoyager conditions each new hypothesis on what has already been attempted, preventing redundancy. It generates code, runs it, interprets outputs via a vision-language model, and updates the exploration plan accordingly.
⚡ Outperforms GPT-4o baseline by 16% (micro-averaged, p<0.01) and 19.33% (macro-averaged, p<0.001) on CellBench - 50 published scRNA-seq studies, 483 analyses. Discovered CD8+ T cells in COVID-19 are more primed for pyroptosis and a link between transcriptional noise and aging in the brain’s subventricular zone - both absent from the original papers.
🔬 Applications & Insights
1️⃣ Hypothesis Generation from Published Data
Proposes novel analytical directions beyond those in the original paper - without new experiments or new data.
2️⃣ COVID-19 Immunology
Found CD8+ T cells in COVID-19 patients are more primed for pyroptosis - a mechanistic insight absent from the original study.
3️⃣ Brain Aging
Discovered a link between transcriptional noise and aging in the subventricular zone, validated as novel by the original authors.
4️⃣ Collaborative Research
Acceleration Ingests what has been done, explores what hasn’t, and produces interpretable reports for expert review.
💡 Why This Is Cool
Most scRNA-seq datasets have more biology in them than the original analysis captured. CellVoyager treats that gap as an opportunity. The benchmark measures whether the agent predicts what a paper actually analysed - a much harder standard than held-out prediction tasks.
📖 Read the paper
💻 GitHub
MacroGuide: Topological guidance for macrocycle generation
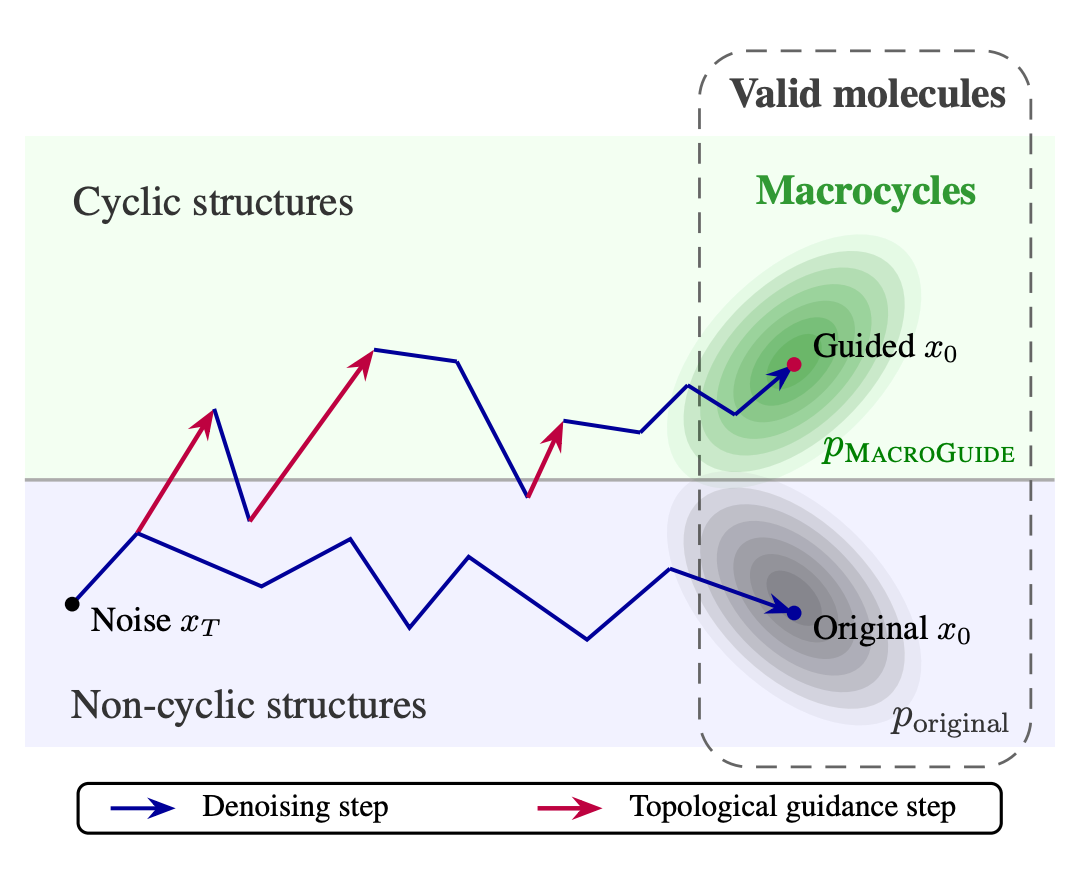
🔬 Macrocycles offer superior selectivity against difficult drug targets, but generative models almost never produce them. The core problem is topological: ring closure is a global structural constraint that local generative approaches can’t enforce.
University of Oxford, AITHYRA, ENS Ulm, and TU Wien introduce MacroGuide, a training-free diffusion guidance mechanism that steers any pretrained molecular generative model toward macrocycles using Persistent Homology.
🧬 At each denoising step, MacroGuide constructs a Vietoris-Rips complex from atomic positions and computes gradients from three topological objectives: cycle size (H1 death), cycle connectivity (H1 birth), and molecule connectivity (H0 death). These steer the score function toward ring-forming structures without modifying the base model.
⚡ Macrocycle generation rate from 1% to 99% on pretrained diffusion models. Matches or exceeds SOTA on chemical validity, diversity, PoseBusters checks, and pharmacophore satisfaction. Demonstrated in unconditional and protein pocket-conditioned settings including bicyclic structures.
🔬 Applications & Insights
1️⃣ De Novo Macrocycle Design
Not limited to cyclic peptides or linear scaffolds - enables arbitrary ring architectures without requiring known linear equivalents.
2️⃣ Plug-and-Play Compatibility
Training-free. Plugs into any pretrained diffusion model without retraining or fine-tuning.
3️⃣ Structure-Based Drug Design
Generates macrocycles conditioned on protein binding pockets, including bicyclic structures, for structure-based drug design campaigns.
4️⃣ Difficult Target Access
Macrocycles bind protein surfaces and allosteric sites unreachable by small molecules. MacroGuide makes these architectures designable at scale.
💡 Why This Is Cool
Macrocycles hit targets that small molecules can’t - but designing them required peptide chemistry or handcrafted scaffolds. MacroGuide removes both constraints as a plug-in for any diffusion-based molecular generation workflow. No retraining, no approximation.
📖 Read the paper
BioReason-Pro: Multimodal biological reasoning for protein function prediction
🔬 Standard protein function prediction treats GO annotation as classification - it gives a label but not a reason. Expert biologists reason across sequence, structure, domains, evolution, and interaction networks. BioReason-Pro was built to do the same.
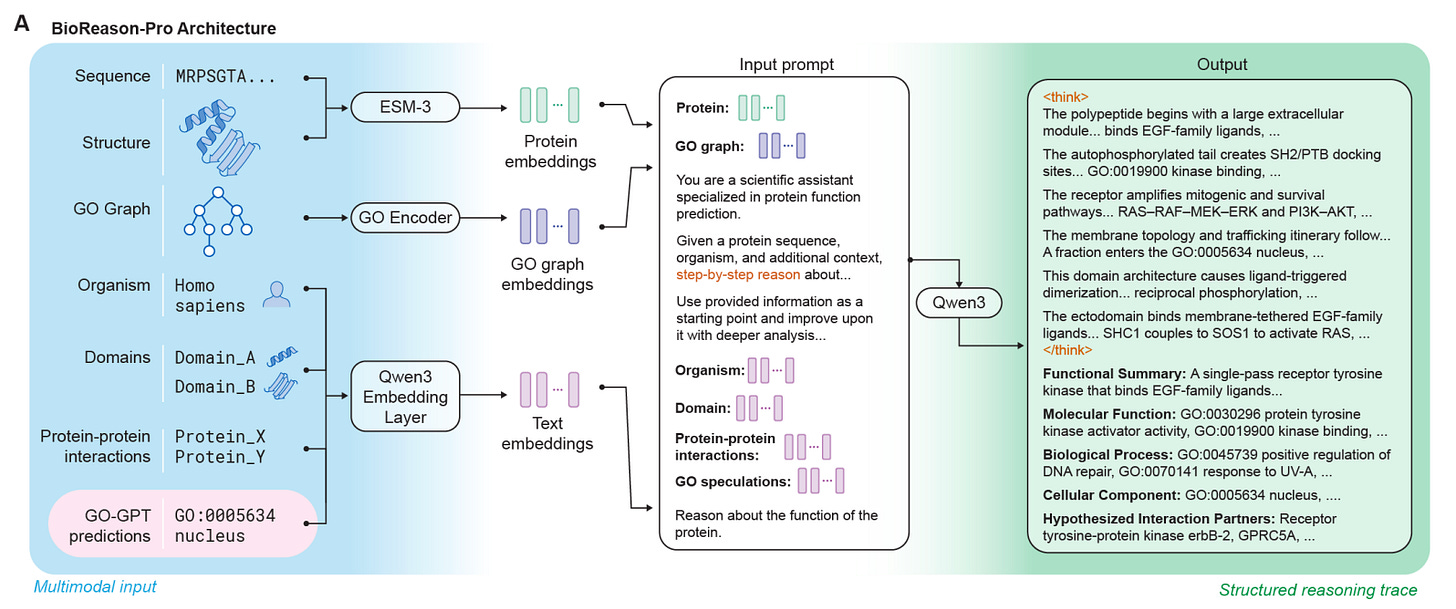
The Arc Institute, University of Toronto, Vector Institute, and Stanford built BioReason-Pro, the first multimodal reasoning LLM for protein function prediction, generating structured step-by-step biological reasoning traces.
🧬 BioReason-Pro integrates ESM3 residue embeddings, GO graph structure, STRING protein interactions, and InterPro domain annotations. GO-GPT, an autoregressive transformer, provides sequential GO term predictions as context. Fine-tuned on 133K+ proteins via SFT on GPT-5-generated reasoning traces, then optimised with RL using GO prediction accuracy as reward.
⚡ GO-GPT achieves F_max = 0.70 (best-of-10), outperforming InterLabelGO+ and ProtBoost. Human protein experts preferred BioReason-Pro over ground truth UniProt annotations in 79% of cases. Attention for DNA-binding proteins aligns to exact binding residues (AUROC 0.81, 2.8x fold-enrichment) - without structural input.
🔬 Applications & Insights
1️⃣ Protein Annotation at Scale
Applied to 240,000+ proteins including the Human Protein Atlas, covering the vast majority with no experimental annotations.
2️⃣ Binding Partner Prediction
Zero-shot identification with attention aligning to cryo-EM-resolved contact residues - structural validation without structural input.
3️⃣ Beyond Homology
Integrates multiple evidence types to override misleading superfamily annotations, beyond what homology transfer alone achieves.
4️⃣ Understudied Proteins
Robust on proteins with low training similarity - relevant for viral proteins, rare organisms, and novel therapeutic targets.
💡 Why This Is Cool
Most protein function tools give you a label. BioReason-Pro gives you a reason. Human experts preferred its annotations over UniProt ground truth in 79% of cases - the model’s reasoning is considered more credible than the existing gold standard. That’s a meaningful bar.
📖 Read the paper
💻 GitHub
🗓️ Events & Competitions
The best competitions, hackathons, and community challenges in AI x life sciences, curated weekly. Know something worth featuring? Reply and let us know.
Check this section next week for a recap with the organisers of BioHackathon Edinburgh 2026!
More upcoming events:
BioHackathon Europe 2026 | November 9-13, Barcelona
ELIXIR’s annual international bioinformatics hackathon, running since 2018. 160+ participants, five days of collaborative coding on open bioinformatics infrastructure and tools. The call for project proposals opens March 16 and closes April 15 - so if you want to lead a project, that’s your window.
Thanks for reading!
💬 Get involved
We’re always looking to grow our community. If you’d like to get involved, contribute ideas or share something you’re building, fill out this form or reach out to me directly.
Connect With Us
Have questions or suggestions? We'd love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website