Stanford's dEVA, McMaster's SyntheMol-RL, and SNU's Expression Rescue
Kiin Bio's Weekly Insights
Welcome back to your weekly dose of AI news for Life Science!
Keeping up with AI x life science news can get exhausting.
It’s scattered across LinkedIn, X, Substack, arXiv, Slack, newsletters... and you still somehow miss the things that actually matter. Too much noise, not enough signal.
We’re building something to fix that: a smarter, more powerful way to stay on top of what’s actually relevant to you.
But we want to build it with you, not just for you. Take 2 minutes to tell us what’s missing. What you share will directly shape what we build, and you’ll be the first to benefit from it.
🇺🇸 We’re heading to Bio-IT World in Boston, May 19-21.
Our CEO Filippo and CTO Bogdan will be there and would love to meet anyone thinking about:
How AI is actually changing preclinical workflows (not just the hype)
Why drug discovery is a systems problem, not just a science one
What it takes to go from 5-year timelines to something radically faster
No pitch, just good conversation. If any of that’s on your mind, reach out - we’ll find a time to grab a coffee.
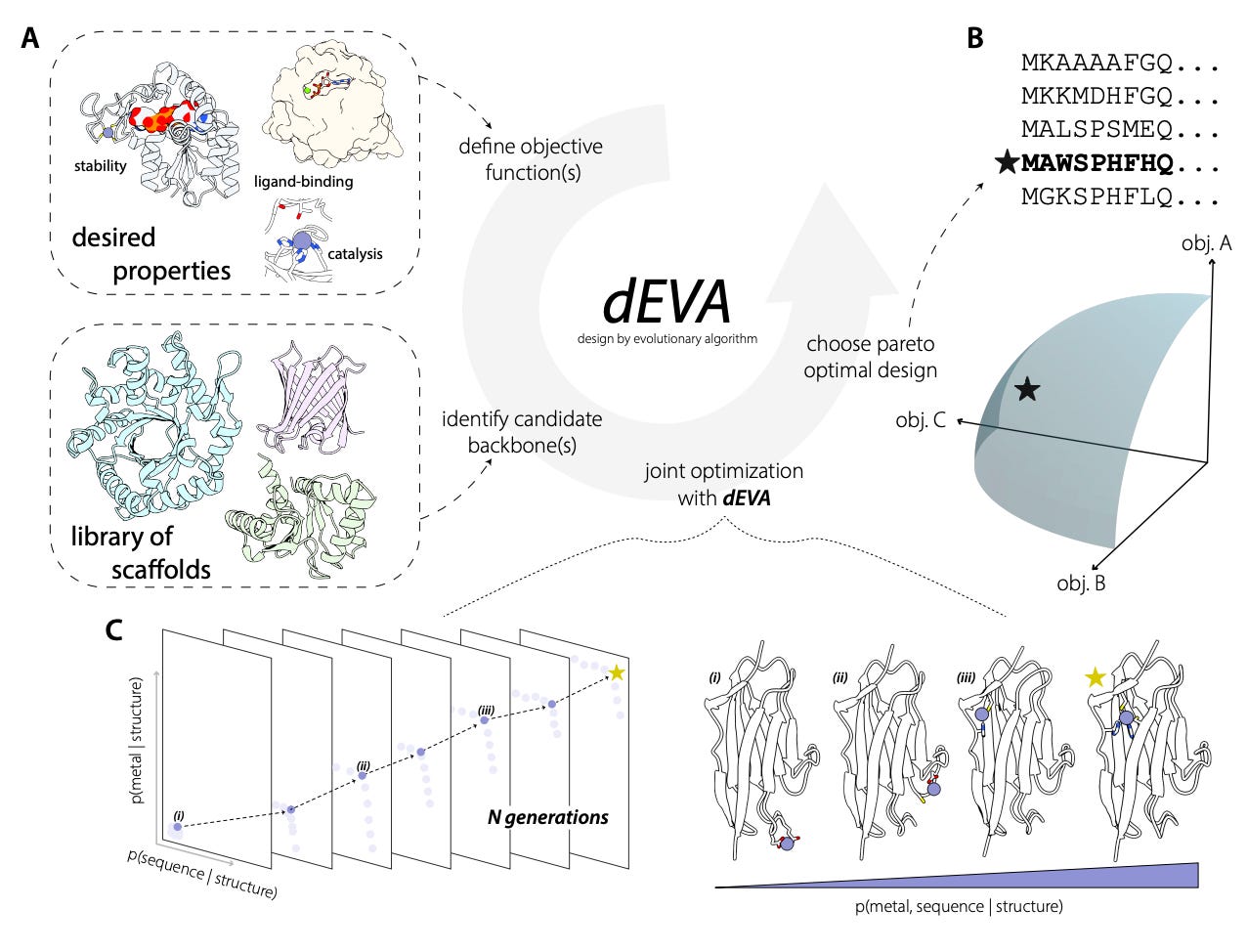
dEVA: Zero-Shot Design of a De Novo Metalloenzyme
🔬 Designing functional enzymes from scratch remains one of the hardest challenges in protein design. Previous approaches relied on borrowing catalytic motifs from nature, but optimising for structure alone does not guarantee catalytic competence. Efficient catalysis requires a precise balance of chemical, geometric, and electrostatic criteria that existing methods struggle to jointly satisfy.
Gina El Nesr and colleagues at Stanford present dEVA (design by EVolutionary Algorithm), a multi-objective framework built on NSGA-II that simultaneously optimises multiple design objectives, enriching for candidates where all criteria are mutually compatible rather than traded off against one another. Using LigandMPNN for sequence design and Metal3D for metal site prediction, dEVA iterates mutations across generations, converging on Pareto-optimal solutions.
🧬 Their best design, desB, achieves catalytic efficiency of 1,500 M⁻¹s⁻¹ and rate enhancement of 3x10¹³ without directed evolution. Analysis of PDB zinc sites revealed around 10% had zero coordinating ligands and over 54% had two or fewer residues, many being crystallisation artefacts. They trained Metal3D-Clean and Metal3D-Cat on curated data to address this.
⚡ desB hydrolyses both phosphomonoesters (uncatalysed half-life >500,000 years) and phosphodiesters (>13 million years). This promiscuity mirrors early enzyme evolution, opening the door to engineering specificity from a designed starting point.
🔬 Applications and Insights
1️⃣ Zero-Shot Functional Enzyme
Design dEVA designed a catalytically active metalloenzyme without natural templates or evolutionary information. No directed evolution was needed to achieve function.
2️⃣ Multi-Objective Optimisation Over Single-Score Ranking
By treating design as population-based evolutionary search across multiple objectives, dEVA avoids the compromises of single-score optimisation or sequential filtering.
3️⃣ Training Data Quality Matters
Curating PDB metal sites and retraining Metal3D on catalytically relevant examples was essential. Garbage in, garbage out applies to structural prediction too.
4️⃣ A Platform for Designed Catalysis
The promiscuous activity of desB mirrors early enzyme evolution, providing a starting point from which specificity can be engineered.
💡 Why This Is Cool
This is the first de novo enzyme designed without borrowing from nature that matches natural catalytic efficiency. The rate enhancement of 3x10¹³ is the highest for any de novo designed hydrolase. dEVA shows functional catalytic sites can emerge computationally from first principles.
📄 Read the paper
💻 Try the code
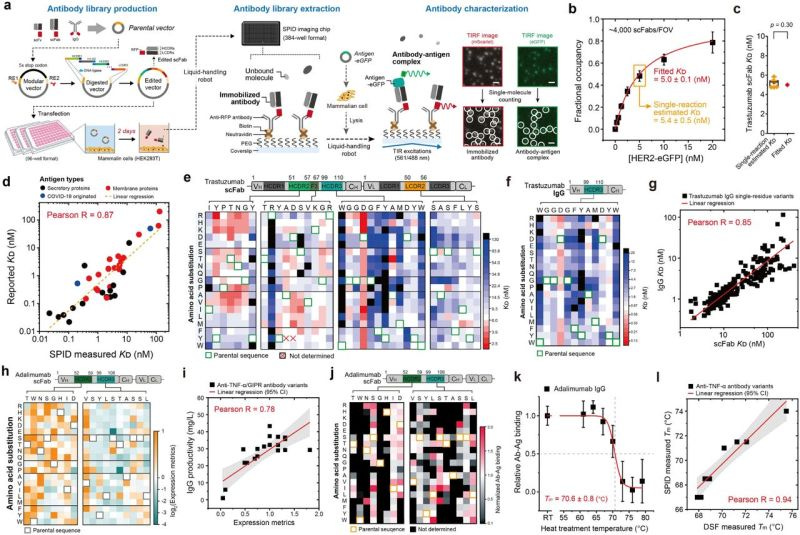
Expression Rescue: Structure-Guided Recovery of Antibody Productivity
🔬 High-affinity antibody variants often fail in production because of poor cellular expression. Researchers at Seoul National University combined AlphaFold3 and ProteinMPNN into a rescue workflow that identifies sequence-structure mismatches in CDR residues and corrects them, often with a single substitution, restoring expression while preserving binding affinity.
🧬 Affinity and expression are largely independent, meaning high-affinity, low-productivity (HALP) clones are not failures. They can be rescued.
⚡ ProteinMPNN scores correlate strongly with cellular productivity across around 9,500 variants, revealing that expression is governed by how well CDR sequences fit their structural context, not just their biochemical properties.
🔬 Applications and Insights
1️⃣ Rescuing High-Affinity Failures
Across 14 diverse HALP antibodies, single-residue substitutions restored expression in 11 cases, while preserving 80% or more of original binding affinity. This reframes failed candidates as recoverable assets rather than endpoints.
2️⃣ Sequence-Structure Compatibility as a Predictor
ProteinMPNN scores serve as a reliable proxy for expression, providing a computationally cheap filter before expensive experimental validation.
3️⃣ Minimal Edits, Maximal Impact
Rescue often required only one mutation, yielding up to 4-fold improvements in expression. These substitutions typically stabilise interactions within CDRs or between CDRs and the antibody framework.
4️⃣ Decoupling Affinity and Developability
Because productivity and affinity landscapes are independent, expression can be improved without compromising binding, solving a long-standing trade-off in antibody engineering.
💡 Why This Is Cool
Antibody design has long focused on finding better binders, but binding is only half the story. By reframing expression as a structural compatibility problem, failed candidates become fixable rather than disposable. This turns antibody engineering from a filtering step into a repair-and-optimise loop, expanding the usable therapeutic space.
📄 Read the paper
💻 Try the code
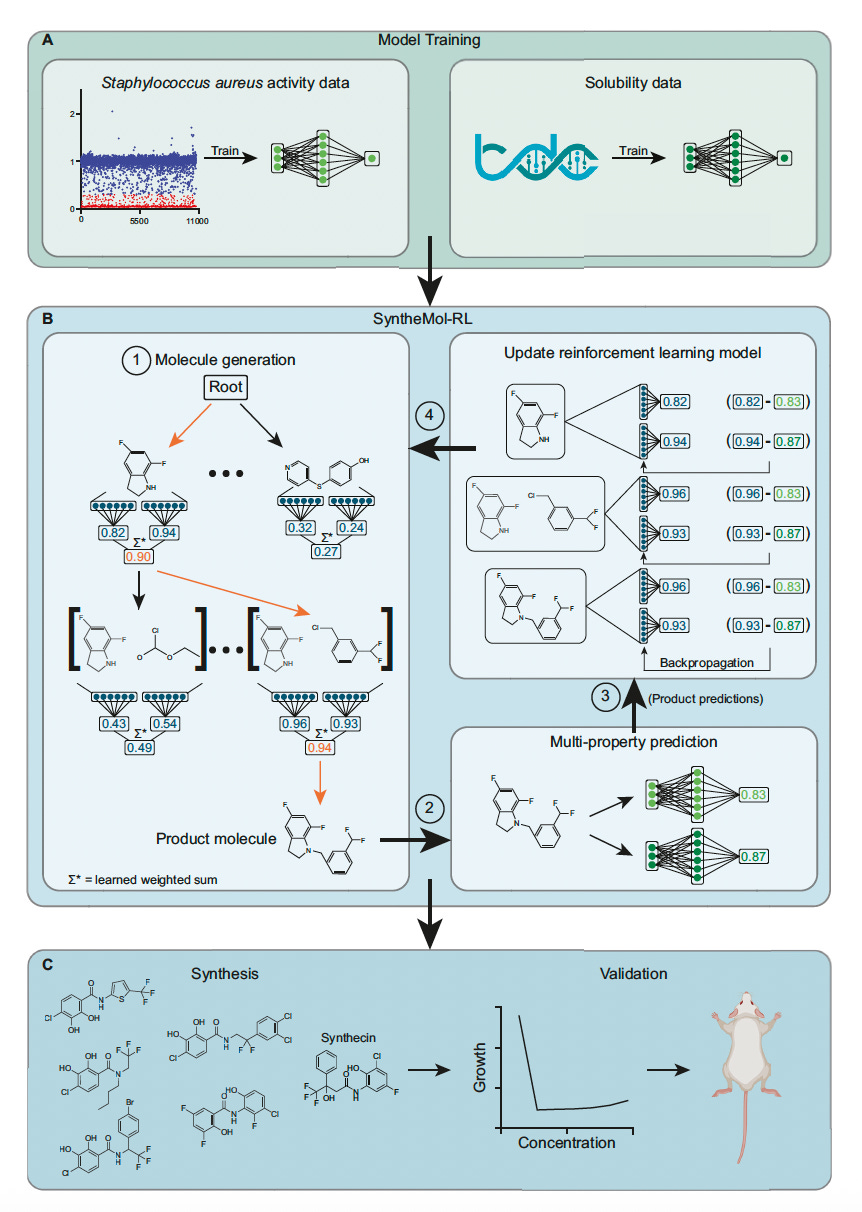
SyntheMol-RL: Reinforcement Learning for Designing Easily Synthesizable Antibiotics
🔬 Generative AI can propose drug candidates, but most fail at the same hurdle: they cannot be synthesised efficiently. Molecules that look promising on screen often require impractical chemistry to produce. SyntheMol-RL from Stanford and McMaster University uses reinforcement learning to navigate a chemical space of 46 billion synthesisable compounds, optimising for both antibacterial activity and aqueous solubility simultaneously.
🧬 Built on real chemical building blocks and validated reaction templates, the model generates molecules with guaranteed synthetic routes. This is not theoretical synthesisability. Every output comes with a concrete pathway from purchasable reagents.
⚡ 79 novel compounds were synthesised and tested. 13 showed potent in vitro activity against Staphylococcus aureus. Seven passed structural novelty filters against known antibiotics. One compound, synthecin, demonstrated efficacy in a murine wound infection model of methicillin-resistant S. aureus (MRSA).
🔬 Applications and Insights
1️⃣ From Screen to Syringe
Unlike virtual screening, SyntheMol-RL generates molecules with built-in synthesis plans, removing the bottleneck between computational hits and experimental validation.
2️⃣ Multi-Parameter Optimisation
Reinforcement learning enables simultaneous tuning of activity, solubility, and synthesisability rather than optimising one property at the cost of others.
3️⃣ In Vivo Validation
Synthecin’s efficacy in a wound infection model moves AI-designed antibiotics beyond petri dish activity into preclinical relevance.
4️⃣ Generalisable Framework
The architecture is target-agnostic. Swap the reward function and the same framework applies across therapeutic domains, not just antibiotics.
💡 Why This Is Cool
Most generative models for drug discovery propose molecules that cannot be made, or can be made but do not work. SyntheMol-RL closes both gaps: it only proposes what chemistry can deliver, and it validated a compound through to animal models. Going from 46 billion possibilities to a single molecule treating MRSA-infected wounds in mice is the full loop from generative AI to preclinical candidate.
📄 Read the paper
💻 Try the code
📬 Newsletter Shout-Out
This week we're shouting out Building in BioAI, a monthly newsletter from Joe:
Building in BioAI is a monthly newsletter for those operating in, or interested in, the AI-enabled biology space. That’s founders, technical leaders, and individual contributors working within areas like therapeutics, diagnostics, and tooling.
Joe’s roundup centres on observations from within the space, including analysis of how teams are structuring themselves, what’s changing in hiring, where funding is landing, what headlines mean for growth, and how BioAI companies are thinking about commercialising what they’re building.
Each edition pulls from ongoing conversations with people doing the work day-to-day, as well as his own take on what’s hit headlines that month.
Joe recruits in this space day-to-day, and so often speaks from that vantage point. He spends most of his time inside these teams, hiring for them, speaking with founders and senior talent across the market. The aim isn’t to overstate where things are going, but to give a clear picture of what’s actually happening and why it matters if you’re hiring or looking to commercialise in BioAI.
🗓️ Events & Competitions
The best competitions, hackathons, and community challenges in AI x life sciences, curated weekly. Know something worth featuring? Reply and let us know.
More upcoming events:
BioHackathon Europe 2026 | November 9-13, Barcelona
ELIXIR’s annual international bioinformatics hackathon, running since 2018. 160+ participants, five days of collaborative coding on open bioinformatics infrastructure and tools. The call for project proposals opens March 16 and closes April 15 - so if you want to lead a project, that’s your window.
Thanks for reading!
💬 Get involved
We’re always looking to grow our community. If you’d like to get involved, contribute ideas or share something you’re building, fill out this form or reach out to me directly.
Connect With Us
Have questions or suggestions? We'd love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website