Stanford's Eubiota, NVIDIA's Fold-CP, and Xaira's X-Cell
Kiin Bio's Weekly Insights
Welcome back to your weekly dose of AI news for Life Science!
Keeping up with AI x life science news can get exhausting.
It’s scattered across LinkedIn, X, Substack, arXiv, Slack, newsletters... and you still somehow miss the things that actually matter. Too much noise, not enough signal.
We’re building something to fix that: a smarter, more powerful way to stay on top of what’s actually relevant to you.
But we want to build it with you, not just for you. Take 2 minutes to tell us what’s missing. What you share will directly shape what we build, and you’ll be the first to benefit from it.
Eubiota: Modular agentic AI for autonomous discovery in the gut microbiome
🔬 Microbiome research generates vast genomic, metabolic, and clinical data, but connecting it to mechanistic discoveries requires navigating thousands of tools, databases, and experimental protocols. Most AI systems handle isolated steps - none handle the full scientific workflow.
Stanford University and Chan Zuckerberg Biohub built Eubiota, an open-source agentic AI framework with specialised agents for planning, execution, verification, and synthesis, connected through shared memory and reinforcement-learned reasoning.
🧬 Eubiota coordinates domain-specific tools across the microbiome research workflow: gene screening, metabolite analysis, experimental design, and literature synthesis. Reinforcement learning optimises multi-turn reasoning, allowing the system to refine hypotheses across iterative experimental cycles.
⚡ 87.7% accuracy on microbiome benchmarks, exceeding GPT-5.1 by 10.4%. Screened nearly 2,000 bacterial genes to identify the uvr-ruv DNA repair axis as a fitness determinant under inflammatory stress. Designed a four-strain consortium that reduced colitis severity in mice. Generated a commensal-sparing antibiotic cocktail and discovered diet-associated metabolites suppressing NF-kB signalling.
🔬 Applications & Insights
1️⃣ Autonomous Gene Screening
Screened ~2,000 bacterial genes to identify mechanisms driving fitness under gut inflammation - a scale and speed not achievable with manual experimental design.
2️⃣ Therapeutic Consortium
Design Designed a four-strain bacterial consortium that reduced colitis severity in a mouse model, demonstrating the framework’s ability to move from discovery to experimental validation.
3️⃣ Antibiotic Development
Generated a commensal-sparing antibiotic cocktail - targeting pathogens while preserving the healthy microbiome, a longstanding challenge in antimicrobial therapy.
4️⃣ Metabolite-Immune Crosstalk
Identified diet-associated metabolites that suppress NF-kB signalling, opening new angles for dietary and metabolic interventions in inflammatory disease.
💡 Why This Is Cool
Eubiota changes what’s possible in microbiome research by removing the coordination bottleneck. The most valuable finding here isn’t any single discovery - it’s that an AI system can now run the iterative hypothesis-experiment-synthesis loop that normally takes a large research team years. That’s a different category of tool.
📖 Read the paper.
💻 Try the code.
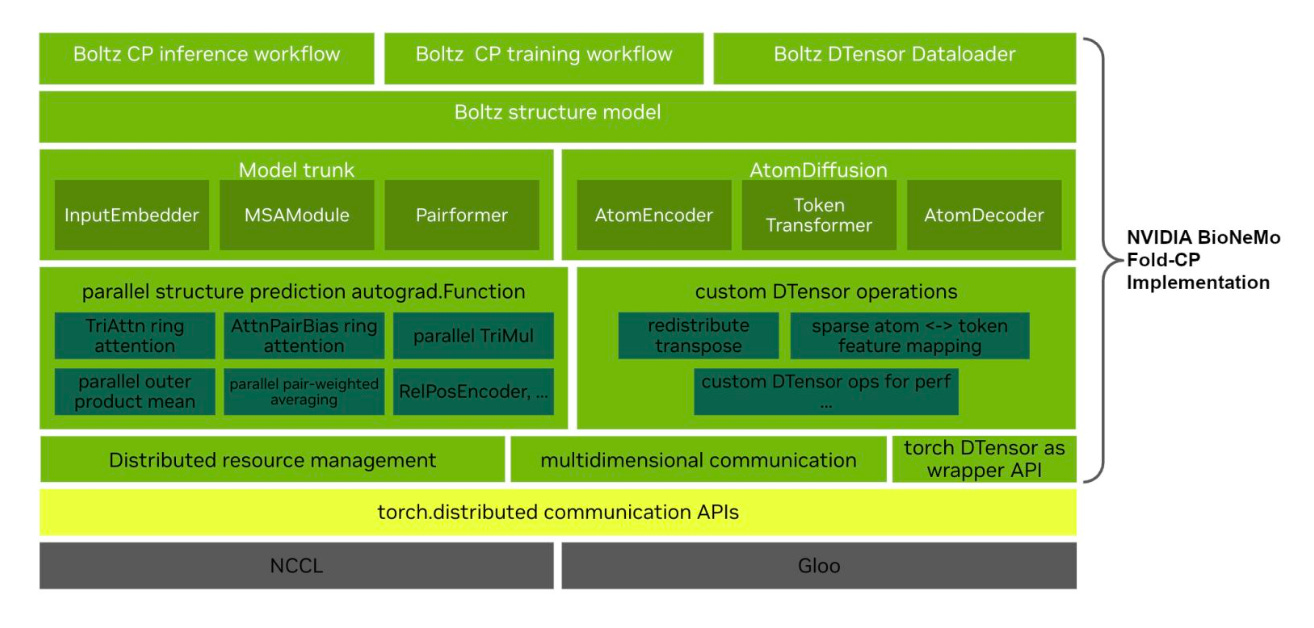
Fold-CP: A context parallelism framework for biomolecular modelling
🔬 AlphaFold-style models use a pairwise representation that scales quadratically with sequence length, capping useful inference at a few thousand residues on a single GPU. Over 70% of characterised mammalian protein complexes in the CORUM database exceed this limit, leaving most disease-relevant assemblies structurally inaccessible.
NVIDIA BioNeMo Fold-CP distributes the inference and training of co-folding models across multiple GPUs, using the open-source Boltz architecture as a reference.
🧬 A 2D device mesh tiles the N×N pair representation across GPUs so per-device memory scales as O(N²/P). Custom distributed algorithms handle triangle attention, triangle multiplication, and window-batched local attention without cropping, chunking, or approximation.
⚡ Structure prediction of assemblies exceeding 30,000 residues on 64 NVIDIA B300 GPUs. Accuracy parity with single-device baselines (R=0.97, median lDDT difference 0.0007). Scored 93% of the CORUM mammalian complex database, up from <30% accessible on standard models.
🔬 Applications & Insights
1️⃣ Mega-Complex Structure Prediction
Assemblies exceeding 30,000 residues predicted on 64 NVIDIA B300 GPUs. Over 70% of CORUM mammalian complexes now structurally accessible, up from <30% on standard single-GPU models.
2️⃣ Disease-Relevant Targets
PI4KA, a 3,605-residue complex linked to inflammatory bowel disease, predicted with TM-score 0.83. Previously inaccessible to co-folding models on standard hardware.
3️⃣ Accuracy at Scale
Prediction quality matches single-device baselines (R=0.97, median lDDT difference 0.0007), confirming parallelisation does not degrade structure quality.
4️⃣ Memory Scaling without Approximation O(N²/P) memory scaling across P GPUs. No cropping, chunking, or approximation. Custom distributed triangle attention and multiplication preserve full-model accuracy.
💡 Why This Is Cool
The bottleneck in structural biology has shifted from sequence to scale. Fold-CP doesn’t approximate around the quadratic memory wall, it removes it. The 93% CORUM coverage figure is striking: most disease-relevant complexes were structurally inaccessible before this. That changes what’s tractable for drug discovery.
📖 Read the paper
💻Try the code.
X-Cell: Scaling causal perturbation prediction across diverse cellular contexts
🔬 Most transcriptomic perturbation models train on a single cell type and fail elsewhere. Xaira Therapeutics built both the dataset and the model to change that: X-Atlas/Pisces, 25.6 million perturbed single-cell transcriptomes across 16 contexts, and X-Cell, a diffusion language model that predicts genome-wide perturbation responses.
🧬 X-Cell refines predictions by iteratively denoising control-to-perturbed transitions, conditioning on five biological priors via cross-attention: STRING, ESM-2, DepMap, GenePT, and CellPairing.
⚡ Up to 5-fold improvement over SOTA (Pearson Delta 0.51 vs STATE’s 0.10). Zero-shot T cell inactivation prediction in Jurkat cells. X-Cell-Ultra at 4.9 billion parameters is the first perturbation model to follow LLM power-law scaling. Zero-shot generalisation to iPSC melanocytes and primary CD4+ T cells.
🔬 Applications & Insights
1️⃣ Drug Target Prioritisation
Predict transcriptomic effects of gene knockouts across diverse cell types, reducing reliance on costly per-indication Perturb-seq screens.
2️⃣ Immunotherapy
Zero-shot prediction of T cell inactivating perturbations - identifying knockouts that shift activated T cells toward a resting state for autoimmune and immunotherapy applications.
3️⃣ Drug Response Prediction
X-Cell transfers from genetic to chemical perturbations (Pearson Delta 0.31 vs STATE’s 0.22 on Tahoe-100M drug data), opening paths to in silico drug screening.
4️⃣ Perturbation Biology Scales Like LLMs
X-Cell-Ultra follows LLM-style power-law scaling - the first evidence that larger models and more data consistently improve perturbation prediction.
💡 Why This Is Cool
Most perturbation models answer one cell type’s question. X-Cell answers across the disease-relevant landscape. The power-law scaling finding matters: it means this approach will keep improving with more data and compute - the same engine that drove LLM breakthroughs.
📖 Read the paper.
💻Try the code.
🗓️ Events & Competitions
The best competitions, hackathons, and community challenges in AI x life sciences, curated weekly. Know something worth featuring? Reply and let us know.
More upcoming events:
BioHackathon Edinburgh 2026 | March 20-22, Edinburgh
Three days at the University of Edinburgh bringing together life scientists, programmers, and industry partners to hack on real biological challenges. Tracks cover academic research (gene regulation, drug discovery, imaging), industry innovation, and a non-coder track for experimental design and project management. Applications are closed, but one to watch if you’re at a UK university for next year.
BioHackathon Europe 2026 | November 9-13, Barcelona
ELIXIR’s annual international bioinformatics hackathon, running since 2018. 160+ participants, five days of collaborative coding on open bioinformatics infrastructure and tools. The call for project proposals opens March 16 and closes April 15 - so if you want to lead a project, that’s your window.
Thanks for reading!
💬 Get involved
We’re always looking to grow our community. If you’d like to get involved, contribute ideas or share something you’re building, fill out this form or reach out to me directly.
Connect With Us
Have questions or suggestions? We'd love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website