Yale's HEIST, A-Alpha Bio's SEPIA, and Harvard's Apollo
Kiin Bio's Weekly Insights
Welcome back to your weekly dose of AI news for Life Science!
Keeping up with AI x life science news can get exhausting.
It’s scattered across LinkedIn, X, Substack, arXiv, Slack, newsletters... and you still somehow miss the things that actually matter. Too much noise, not enough signal.
We’re building something to fix that: a smarter, more powerful way to stay on top of what’s actually relevant to you.
But we want to build it with you, not just for you. Take 2 minutes to tell us what’s missing. What you share will directly shape what we build, and you’ll be the first to benefit from it.
🇺🇸 We’re heading to Bio-IT World in Boston, May 19-21.
Our CEO Filippo and CTO Bogdan will be there and would love to meet anyone thinking about:
How AI is actually changing preclinical workflows (not just the hype)
Why drug discovery is a systems problem, not just a science one
What it takes to go from 5-year timelines to something radically faster
No pitch, just good conversation. If any of that’s on your mind, reach out - we’ll find a time to grab a coffee.
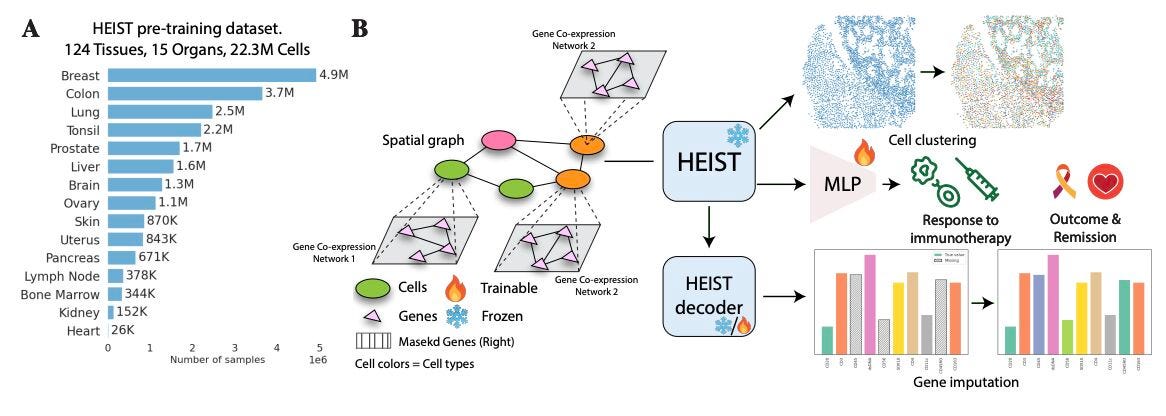
HEIST: A Graph Foundation Model for Spatial Transcriptomics and Proteomics
🔬 Spatial transcriptomics captures gene expression within tissue architecture, but existing models either ignore spatial relationships or flatten each cell into a simple feature vector. They miss the interplay between a cell’s internal gene programmes and its tissue neighbourhood.
HEIST from Yale models tissues as hierarchical graphs. The upper level captures spatial relationships between cells, while each cell is represented by its own gene co-expression network. Cross-level message passing connects the two, letting internal regulation and spatial context inform each other.
🧬 Pretrained on 22.3 million cells from 124 tissues across 15 organs using spatially-aware contrastive learning and masked autoencoding. HEIST uses flexible gene vocabularies rather than fixed gene sets, so it generalises to unseen genes and even spatial proteomics without retraining.
⚡ Unsupervised analysis reveals spatially informed cell subpopulations missed by prior models. Downstream, HEIST achieves state-of-the-art performance in clinical outcome prediction, cell type annotation, and gene imputation across multiple spatial technologies.
🔬 Applications and Insights
1️⃣ Cross-Modal Generalisation
Transfers from transcriptomics to proteomics without retraining, making it applicable across spatial profiling technologies.
2️⃣ Tissue Microenvironment Discovery
The hierarchical design captures spatially defined subpopulations that flat models miss, enabling more nuanced tissue phenotyping.
3️⃣ Clinical Outcome Prediction
Patient-level embeddings from spatial data support tasks like treatment response and survival prediction.
4️⃣ Flexible Gene Vocabularies
By avoiding fixed gene sets, HEIST handles new panels and custom targets without architectural changes.
💡 Why This Is Cool
Most spatial models look at where cells are or what they express. HEIST does both through hierarchical graph modelling. Generalising to proteomics without retraining suggests these representations capture something fundamental about how cells organise within tissues.
📄 Read the paper.
💻 Try the code.
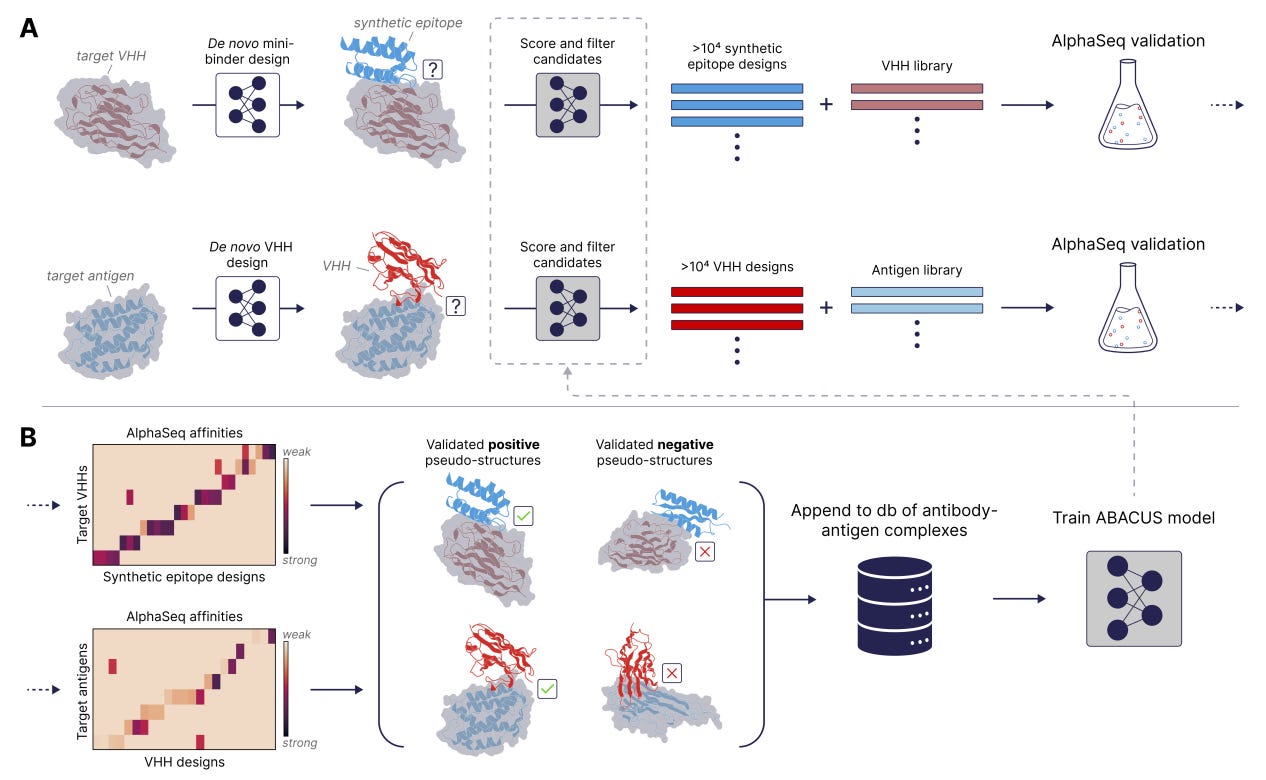
The Synthetic Epitope Atlas: High-Throughput Design and Validation of De Novo Antibody-Antigen Complexes
🔬 ML models for antibody design are held back by a data bottleneck: not enough structural training data linking designed antibodies to validated binding outcomes. Existing datasets are small, biased towards natural antibodies, and lack systematic off-target measurements.
A-Alpha Bio built SEPIA (Synthetic Epitope Atlas), pairing over 26 million on-target and off-target binding measurements with computationally designed VHH-antigen structures. Using their AlphaSeq yeast-based platform, they measured binding affinities and specificity across thousands of de novo synthetic epitope proteins designed to bind VHH nanobodies.
🧬 Each designed VHH-SEP pair comes with both structural predictions and experimental binding data, so models can learn what makes a designed complex actually bind versus what only looks good computationally.
⚡ Across thousands of variants, SEPIA validates strong specificity and provides the negative data most antibody datasets lack. Positive and negative measurements at this scale give ML models a clearer signal for learning specificity, not just affinity.
🔬 Applications and Insights
1️⃣ Training Data for Antibody ML
26 million measurements paired with designed structures create a purpose-built resource for next-generation antibody design models.
2️⃣ Specificity, Not Just Affinity
Systematic off-target measurements let models learn what not to bind, addressing a major blind spot in current datasets.
3️⃣ Closing the Design-Validation Loop
Linking computational designs directly to high-throughput experimental readouts enables rapid iteration on antibody engineering.
4️⃣ Nanobody-Focused Design
VHH nanobodies are increasingly important as therapeutics. A large-scale, VHH-specific dataset accelerates this growing field.
💡 Why This Is Cool
The gap between computational antibody design and experimental reality has always been the data. SEPIA fills it with 26 million purpose-built binding measurements, including both what works and what does not. Models trained on real specificity data at this scale can finally learn to design antibodies that are specific, not just tight binders.
📄 Read the paper.
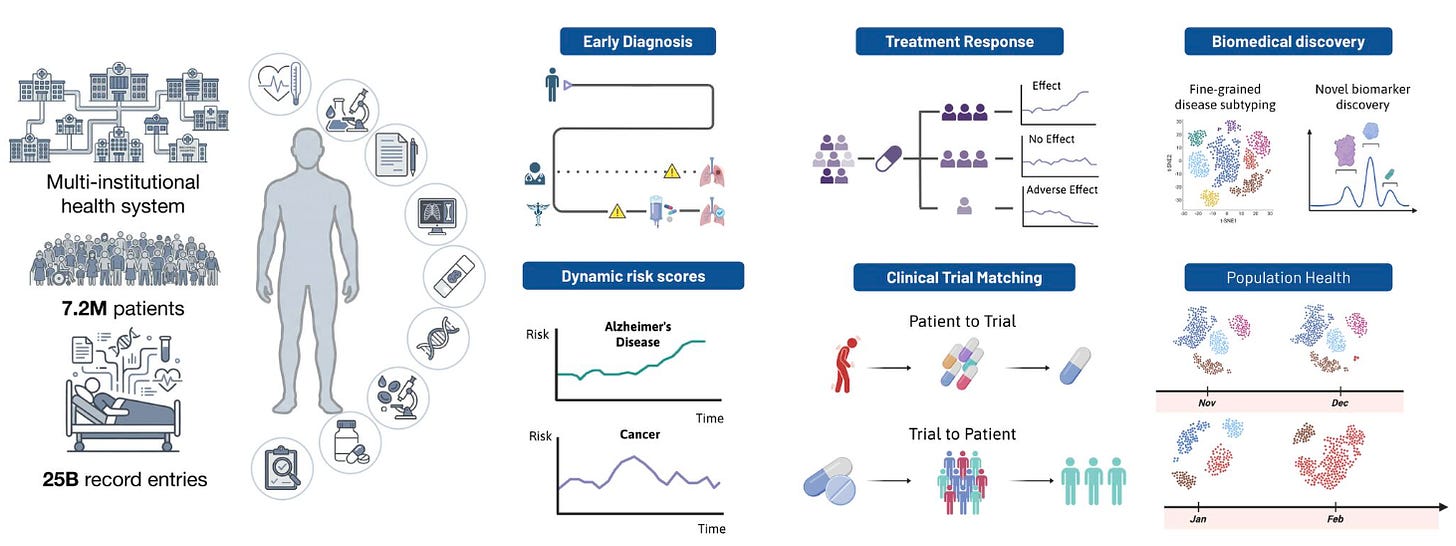
Apollo: A Multimodal Temporal Foundation Model for Virtual Patient Representations
🔬 Modern hospitals generate vast multimodal data across labs, imaging, notes, medications, and procedures, but it sits in disconnected systems. No existing model integrates the full breadth and temporal depth of a clinical record into one unified representation.
Apollo from Harvard Medical School does exactly that. Trained on over 30 years of longitudinal records from Mass General Brigham, it unifies 28 modalities and 12 major specialties into a shared embedding space, building an “atlas of medical concepts.”
🧬 Apollo processes entire care journeys as sequences of structured and unstructured events, compressing them into virtual patient representations. Its vocabulary spans over 100,000 unique medical events alongside clinical images and free-text notes. Feature attribution confirms predictions align with clinically interpretable biomarkers.
⚡ Evaluated across 322 tasks on 1.4 million held-out patients: disease onset prediction up to five years ahead (95 tasks), disease progression (78), treatment response (59), adverse event risk (17), and hospital operations (12). Apollo also functions as a multimodal medical search engine across 61 retrieval tasks.
🔬 Applications and Insights
1️⃣ Five-Year Disease Forecasting
Predicting disease onset years in advance from the full patient record enables proactive intervention rather than reactive care.
2️⃣ Treatment Response Prediction
Drawing on a patient’s complete multimodal history supports more personalised therapy decisions across specialties.
3️⃣ Multimodal Medical Search
Text and image queries against patient embeddings create a clinical search engine for cohort identification and case matching.
4️⃣ Interpretable Predictions
Feature attribution shows outputs align with known biomarkers, bridging AI predictions and clinical reasoning.
💡 Why This Is Cool
This is the first model to compress decades of multimodal clinical data into unified patient embeddings at hospital system scale. Moving from narrow, task-specific clinical models to holistic representations that predict disease, treatment response, and adverse events from one embedding is a fundamental shift for clinical AI.
📄 Read the paper.
🗓️ Events & Competitions
The best competitions, hackathons, and community challenges in AI x life sciences, curated weekly. Know something worth featuring? Reply and let us know.
📄Recap post: BIOMICS Hackathon | Feb 23-25
The BIOMICS hackathon at EMBL-EBI brought together computational biologists and software engineers from Portugal, Spain, Germany, and the UK for three days of building. Five challenge tracks covered everything from statistical tools to building software for a brand new microscopy technique from scratch.
Every team built something visual, reflecting a shift away from command-line-only workflows. This was also one of the first hackathons where AI coding agents like Claude Code were widely used across teams, and the difference in what could be achieved in three days was significant. One participant described these tools as an “exoskeleton” that amplifies existing ability.
BIOMICS is a EU Horizon-funded project twinning GIMM Lisbon with EMBL-EBI, CRG Barcelona, and ETH Zurich to strengthen biomedical data science training and collaboration. More events are planned throughout the year.
More upcoming events:
BioHackathon Europe 2026 | November 9-13, Barcelona
ELIXIR’s annual international bioinformatics hackathon, running since 2018. 160+ participants, five days of collaborative coding on open bioinformatics infrastructure and tools. The call for project proposals opens March 16 and closes April 15 - so if you want to lead a project, that’s your window.
Thanks for reading!
💬 Get involved
We’re always looking to grow our community. If you’d like to get involved, contribute ideas or share something you’re building, fill out this form or reach out to me directly.
Connect With Us
Have questions or suggestions? We'd love to hear from you!
📧 Email Us | 📲 Follow on LinkedIn | 🌐 Visit Our Website